Acinetobacter baumannii is characterized by its tendency to acquire resistance to multiple classes of antimicrobial agents, including carbapenems [Reference Peleg, Seifert and Paterson1]. This resistance has been linked to the presence of integrons and other genetic elements such as the insertion sequence ISAba1. Integrons contain gene cassettes that carry resistance determinants and have been implicated in several outbreaks of opportunist species, particularly A. baumannii, in hospitals [Reference Mazel2]. Five classes of integrons have been described based on the sequence of their intI genes, and class 1 is by far the most prevalent in A. baumannii isolates [Reference Mazel2, Reference Turton3]. The ISAba1 element has been found in association with carbapenem-resistance genes bla OXA-51-like, bla OXA-23-like, bla OXA-58-like and AmpC cephalosporinase. ISAba1 appears to provide a promoter sequence which results in the overexpression of these resistance genes, as well as modulating the mobility of OXA-type genes [Reference Peleg, Seifert and Paterson1, Reference Turton4].
In early 2007, in Porto Alegre, southern Brazil, many hospitals simultaneously reported infections due to multidrug resistant A. baumannii. The increased infection rates persisted for at least 12 months and four intensive care units, in different hospitals, were closed due to this outbreak. A previous study of our group characterized 239 isolates of carbapenem-resistant (meropenem and/or imipenem) A. baumannii obtained from patients from five hospitals between July 2007 to June 2008 in Porto Alegre, and identified 14 distinct clonal groups by molecular techniques [Reference Martins5]. From that study we selected a total of 41 carbapenem-resistant isolates as representatives of each clonal group, including at least one isolate from the five hospitals, to evaluate the influence of mobile genetic elements in resistance to carbapenems. We also included 17 carbapenem-susceptible A. baumannii resulting in a total of 58 isolates. The isolates were identified using an automated system (Vitek, bioMérieux, France) and/or standard phenotypic methods performed in the clinical microbiology laboratory of each hospital and species-specific PCR for the bla OXA-51-like gene [Reference Turton6].
Minimal inhibitory concentrations (MICs) were determined by broth microdilution and/or the M.I.C.EvaluatorsTM method (Oxoid, UK) to the following antimicrobial agents: ceftazidime (Novafarma, Brazil), ampicillin/sulbactam (Eurofarma, Brazil), polymyxin B (Eurofarma), imipenem (ABL, Brazil) and meropenem (Eurofarma). Susceptibility results were interpreted according to Clinical and Laboratory Standards Institute guidelines [7]. Pseudomonas aeruginosa ATCC 27853 and Escherichia coli ATCC 25922 were used for quality control.
The presence of bla OXA-51-like, bla OXA-23-like, bla OXA-24-like, bla OXA-58-like and bla OXA-143 was detected by multiplex PCR assay using primers and conditions described previously [Reference Martins5, Reference Woodford8]. The isolates were screened for the presence of class 1 and class 2 integrons by PCR using primers to the integrase gene according to Koeleman et al. [Reference Koeleman9].
The insertion sequence ISAba1 upstream of bla OXA-23-like and bla OXA-51-like was sought with primers ISAba1F and OXA-23-likeR or OXA-51-likeR according to Turton et al. [Reference Turton4]. The detection of ISAba1F/OXA-51-likeR was conducted in 25 μl volumes containing 1 × PCR buffer, 200 μm dNTPs, 1·0 U TopTaq DNA polymerase (Qiagen, Germany), 10 pmol of each primer and 3 μl bacterial lysate. The reaction mix for ISAba1F/OXA23-likeR contained in 25 μl, 1 × PCR buffer, 200 μm dNTPs, 1·0 U TopTaq DNA polymerase (Qiagen), 7 pmol of each primer and 10 μl bacterial lysate. Cycling conditions for both PCR reactions were as described by Segal et al. [Reference Segal, Garny and Elisha10].
IntI1, IntI2, ISAba1/OXA-51 and ISAba1/OXA-23 genes were sequenced on an ABI-PRISM 3100 Genetic Analyzer (ABI Ltd, USA). DNA sequences were analysed using the Basic Local Alignment Search Tool to search Genbank for homologous nucleic acid sequences.
Fisher's exact test was used to compare discrete variables. A P value <0·05 was considered statistically significant. All analyses were performed with SPSS software, version 13.0 (IBM, USA).
All 41 carbapenem-resistant isolates presented a high level of resistance (MIC90 ampicillin/sulbactam, 64/32 μg/ml; MIC90 ceftazidime, >256 μg/ml; MIC90 polymyxin B, 4 μg/ml; MIC90 imipenem, >32 μg/ml; MIC90 meropenem, >32 μg/ml) to the five antibiotics tested in contrast to the 17 carbapenem-susceptible isolates which demonstrated susceptibility to ampicillin-sulbactam and polymyxin B (MIC90 ampicillin-sulbactam, 16/8 μg/ml; MIC90 polymyxin B, 1 μg/ml). Eight (13·8%) isolates showed resistance to polymyxin B (⩾4 μg/ml), including one isolate which was susceptible to carbapenems.
All isolates harboured the bla OXA-51-like gene, characteristic of A. baumannii. Thirty-nine (67·2%) carbapenem-resistant isolates and one carbapenem-susceptible isolate were positive for the bla OXA-23-like gene. Genes for bla OXA-24-like, bla OXA-58-like or bla OXA-143 were not detected in any isolate. ISAba1 was located upstream of the bla OXA-23-like gene in the majority (36/40) of isolates harbouring this gene. All isolates with this association displayed high MICs (MIC90 imipenem, ⩾32 μg/ml; MIC90 meropenem, ⩾32 μg/ml) for both carbapenems tested. In 23 (39·5%) isolates ISAba1 was located upstream of the bla OXA-51-like gene and 14 of these were also positive for ISAba1/bla OXA-23-like, with one isolate having the bla OXA-23-like gene alone. Therefore, eight isolates had the sole association of ISAba1/bla OXA-51-like gene and these were susceptible to at least one of the carbapenems.
Integrase genes were detected in 51 (87·9%) of the 58 isolates analysed and were present in all of the carbapenem-resistant isolates and 10 (62·5%) of the carbapenem-susceptible isolates. Class 2 integrons were the most prevalent (86·2%), 14 (24·1%) isolates had both classes and a single isolate harboured only the class 1 integron. Each of the eight polymyxin-resistant isolates harboured both classes.
According to the molecular characterization of resistance determinants of each isolate, 12 groups were established (Table 1). Five groups (1, 2, 4, 5, 6) included solely carbapenem-susceptible isolates, five groups (8, 9, 10, 11, 12) included carbapenem-resistant isolates only, and isolates from groups 3 and 7 showed variable susceptibility to carbapenems. Although susceptible isolates displayed ISAba1 upstream of bla OXA-51-like, none of these presented this association with the bla OXA-23-like gene (Table 1).
Table 1. Molecular characterization of resistance determinants of carbapenem-resistant strains of A. baumannii
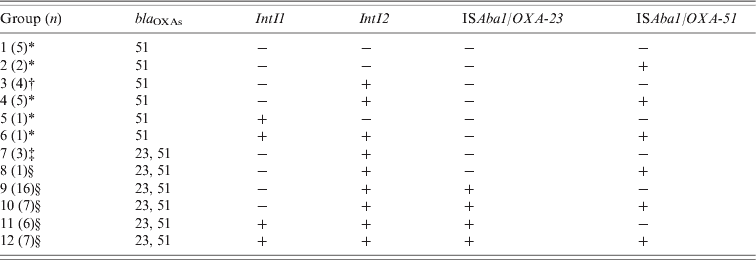
* All isolates of this group were susceptible to carbapenems.
† Two carbapenem-susceptible isolates and two carbapenem-resistant isolates.
‡ One carbapenem-susceptible isolate and two carbapenem-resistant isolates.
§ All isolates of this group were resistant to at least one carbapenem.
Our study included isolates collected over a 1-year period from five major hospitals in Porto Alegre, Brazil. They were selected to be representative of distinct clonal groups, defined earlier by repetitive sequence PCR and pulsed-field gel electrophoresis [Reference Martins5], in order to evaluate the prevalence and dissemination of resistance determinants in different strain clusters. We found that in almost all carbapenem-resistant isolates the bla OXA-23-like gene was accompanied by the ISAba1 element. This was consistent with the premise that the degree of carbapenem resistance in these isolates was accentuated by the presence of promoter sequences provided by ISAba1, leading to expression of the enzyme OXA-23, as previously demonstrated with European strains [Reference Turton4]. The carriage of ISAba1 upstream of bla OXA-23-like in almost all genetically unrelated isolates, obtained in different periods and from different hospitals, indicates that this genetic context is highly transmissible among different strains of carbapenem-resistant A. baumannii.
It is of note that nine (15·5%) isolates with ISAba1 upstream of bla OXA-51-like were susceptible to carbapenems. It might be expected that this combination should confer resistance to the carbapenems, as proposed by Turton et al. [Reference Turton4]. However, our data suggest that the promotion of the bla OXA-51-like without an efficient transcription of the gene is insufficient to confer resistance to carbapenems. These findings were corroborated in a study published by Bratu et al. [Reference Bratu11] in which A. baumannii imipenem-susceptible isolates displayed the ISAba1/bla OXA-51-like association. The lower transcript level of bla OXA-51-like might be due to a dysregulation of the transcription process, or to a disruption of the bla OXA-51-like gene, as already demonstrated [Reference Valenzuela12]. This would justify the fact that, in this study, no statistically significant (P = 0·36) relationship between the presence of ISAba1 upstream of bla OXA-51-like and carbapenem resistance was found. Further studies on gene expression are required to evaluate fully the role of ISAba1 on transcription levels of bla OXA-51-like and bla OXA-23-like genes.
Class 1 integrons are distributed worldwide and are by far the most common in clinical isolates of Gram-negative bacteria, including the genus Acinetobacter [Reference Turton3]. Although class 2 integrons are rare in Acinetobacter species in USA, Europe and Asia [Reference Koeleman9], some studies have demonstrated a high prevalence of this element in Latin American countries, including Brazil [Reference Ramirez13, Reference Fonseca14]. Our study corroborates these data and is the first to report this high prevalence in southern Brazil.
It is important to highlight that isolates belonging to groups 9, 10, 11 and 12 were disseminated among the five hospitals involved in the study and remained for a long period in the hospital setting as previously reported by our group [Reference Martins5]. The ubiquity of class 2 integrons in all carbapenem-resistant isolates underlines their propensity to harbour resistance gene cassettes which can spread and persist in strains of A. baumannii in the hospital environment.
DECLARATION OF INTEREST
None.



