Introduction
The brain contains a large diversity of neuron types, and other cell types like glia, which work together in dense, complex networks to implement behavior, cognition, and emotion. Different neuronal cell types change in different ways in different brain diseases and conditions that affect over a billion people around the world, none of which can be fully cured or ameliorated. These kinds of neuron differ in their shapes and sizes, in what genes they express, in how they are wired, and in how they physiologically affect one another. They compute using a diversity of molecular and physical signals, perhaps most prominently millisecond-timescale electrical signals that are generated in neurons in response to chemical inputs at neuron–neuron connections called synapses, and are integrated toward the firing of millisecond-timescale electrical pulses called action potentials, or spikes, which in turn propagate throughout the complex arbors of neurons, causing release of chemicals at other synapses.
Neural electrical recordings, over the first century of modern neuroscience, enabled the observation of neural electrical activity patterns that are associated with specific behaviors, or with specific brain diseases and conditions, both in humans and in animal model organisms such as mice. But observing a pattern of neural activity in a specific set of neurons during a specific brain state or process does not prove that the neural activity observed plays a causal role in the brain state or process – perhaps the neural activity that is causally involved with the state or process is found elsewhere in the brain, or perhaps one subset of the neural activity observed is more important for the state or process than another subset. Therefore, methods of precisely controlling neural activity, so that its impact on a behavior or a disease can be causally assessed, are necessary. If you could turn on the activity of a specific set of neurons, you could figure out whether they can initiate, sustain, or modulate a given behavioral, cognitive, or emotional process, or a given disease state, or potential therapeutic process. If you could turn off a specific set of neurons, you could figure out whether they are needed for such a state or process. Pharmacological modulation of neurons has been very influential in basic and applied neuroscience, but the effects take place over timescales of seconds to minutes or longer, limited by factors such as the rate of diffusion of drugs into and out of the brain, and furthermore, the effects are felt by multiple types of neuron. Brain stimulation through the delivery of electric fields, magnetic fields, ultrasound, and other forms of energy (e.g., heat), while potentially quite fast, are also nonspecific in their mechanism of stimulation, and thus can affect multiple kinds of neuron within a densely packed neural network.
The toolbox of optogenetics solves this problem. In optogenetics (‘opto’ referring to light, and ‘genetics’ because the toolset is genetically encoded), neuroscientists express genes encoding for microbial rhodopsins, naturally occurring proteins that serve as light-driven ion pumps and channels, found in organisms such as archaea, algae, and bacteria, in genetically targeted neurons so that their electrical activity becomes controllable by light. The word optogenetics is sometimes more broadly used to refer to any genetically encoded tool that enables control of a cellular process with light (Liu and Tucker, Reference Liu and Tucker2017); here we focus on optogenetic control of neural electrical activity via the genetic expression, and light activation, of microbial rhodopsins.
Microbial rhodopsins are seven transmembrane proteins that normally respond to sunlight, capturing solar energy in the form of ion gradients, or serving as simple photosensors for organisms to navigate in their environments. These proteins covalently bind the vitamin A variant all-trans-retinal, which serves as the photosensitive moiety. Upon illumination by light of the appropriate color, all-trans-retinal isomerizes to 13-cis retinal, and the protein then begins a series of rapid conformational changes that result in the fast transport of specific ions from one side of the membrane to the other. These molecules have closed photocycles – the retinal recovers back to the all-trans form in the dark, without the need for other cell types, or enzymes, to facilitate recovery; thus, the molecules can be light-driven over and over again, as self-contained, autonomous units.
In this review, the first half covers a historical perspective on optogenetics. We first discuss early thinking and perspective on the topic, followed by personal reflections on the early days of optogenetic control of neurons. We follow this with a brief summary of the diversity of applications optogenetics has seen in its first decade and a half. We then review the long path of targeted neural activity control technologies that preceded optogenetics, followed by a review of the history of the discovery and characterization of the microbial rhodopsins themselves. The second half of the review delves into the biophysical properties of rhodopsins that make them such great neural control tools. We review the classes and structure of rhodopsins, their photocycles, their photocurrent magnitudes and kinetics, their action spectra (the colors of light that engage them), and their ion selectivity. Our hope here is to provide a comprehensive review at a specific interdisciplinary interface between two fields, which yielded great impact – namely, how the biophysics of these rhodopsins led them to be so useful in neuroscience.
Historical perspective on optogenetics
Concept of optogenetics and early optogenetics experiments
The need for optogenetics, and the specifications desired for the technology to possess, were enunciated long before the technology was actually invented (see Figure 1 for a timeline of some key dates discussed in the first half of this review). Perhaps, Francis Crick was the first to frame the key specifications that the technology should exhibit. As early as 1979, Crick suggested that a technology ‘by which all neurons of just one type could be inactivated, leaving the others more or less unaltered’ would accelerate neuroscience discovery (Crick, Reference Crick1979). Later, in lectures that took place over many years (according to Roger Tsien, who himself explored the topic of optical control of neurons), and that culminated in an influential essay titled ‘The impact of molecular biology on neuroscience’ (Crick, Reference Crick1999), Crick stated that a key need would be ‘to be able to turn the firing of one or more types of neuron on and off in the alert animal in a rapid manner. The ideal signal would be light, probably at an infrared wavelength to allow the light to penetrate far enough. This seems rather far-fetched but it is conceivable that molecular biologists could engineer a particular cell type to be sensitive to light in this way’.
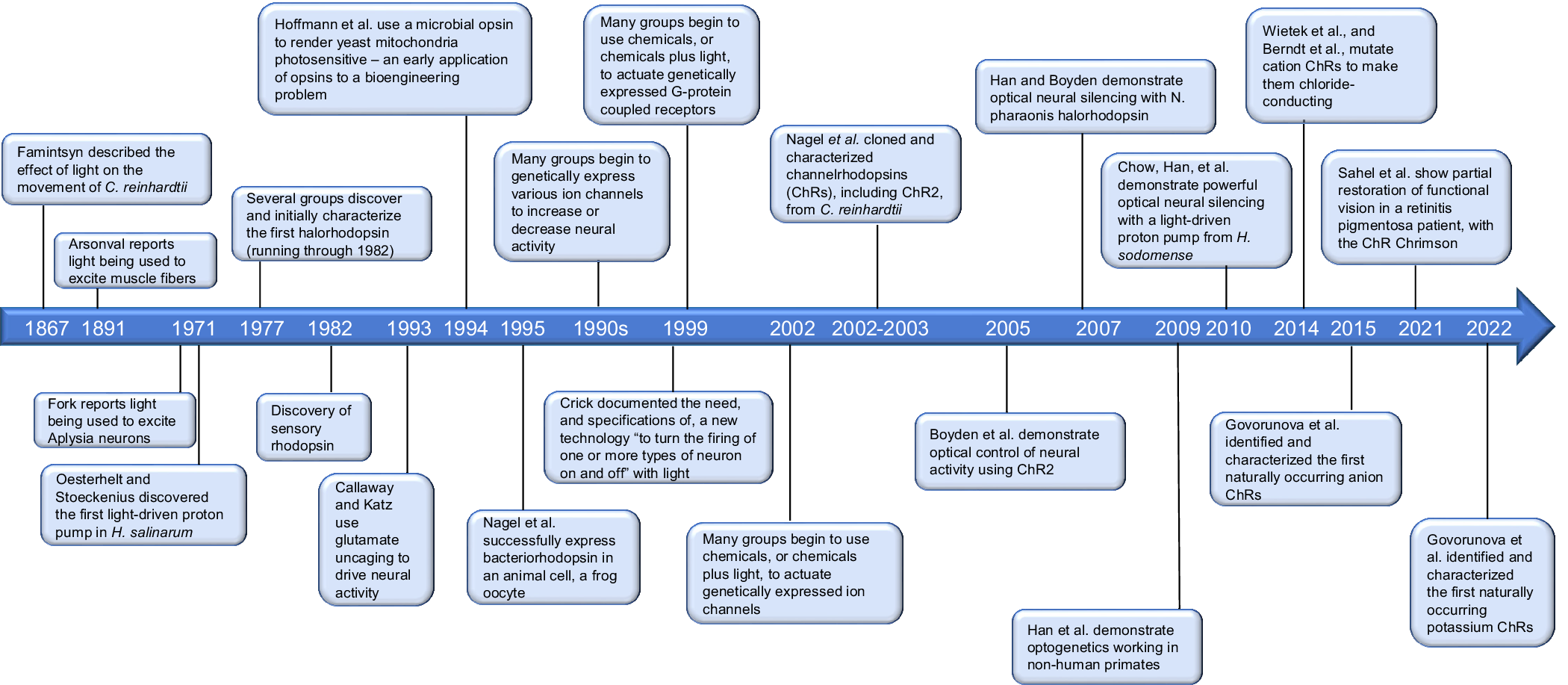
Figure 1. Timeline of key discoveries and innovations in optogenetics.
This was a daunting challenge, as Crick was aware, calling the idea ‘far-fetched’ even in the same breath that he put forth his call to arms: any such technology, to be useful in neuroscience, would have to meet four independent criteria, outlined in Crick’s challenge.
First, the technology should be targetable to a specific neuron ‘type’, and not others, even densely packed neighboring cells that serve functions radically different from those of the targeted type – suggesting the need for genetic targetability of the technology, or something equivalently powerful and easy-to-use.
Second, the technology should be ‘rapid’ enough to keep up with high-speed neural codes, ideally matching the speed of the most fundamental building blocks of brain activity – such as the individual action potential – implying the need for millisecond-timescale temporal precision.
Third, the technology should be easy enough to use, and robust enough, that it could be applied widely in complex neuroscience experiments, even in the delicate, difficult context of the ‘alert animal’ (the simplicity of use of green fluorescent protein (GFP), which needs no chemical supplementation to be used in the awake mammalian brain, comes to mind ( Box 1)).
Box 1. Generalizing Crick’s criteria to other kinds of molecular tool.
It is interesting to think about the generalization of Crick’s criteria, toward general guidelines for creating a molecular technology of great use in biology. Let us consider two examples – GFP and CRISPR – as genetically encoded tools which have had great impact throughout biology. Both of them, curiously enough, do seem to meet generalized forms of Crick’s criteria. First, both are fully genetically encoded and thus can be targeted to different ‘types’ of cell in the living body – GFP, for imaging, and CRISPR, for targeted genome editing. Second, both of them are capable of precision suitable to address the most fundamental building blocks of their respective domain – GFP can be used to visualize individual cells and even molecules, and CRISPR can be used to alter individual genes and even genomic bases. Third, both technologies are very easy to use and robust. GFP needs no chemical supplementation (other than molecular oxygen, which is typically abundant in biological systems) for use in the living cell or body, making it easier to use than earlier genetically targeted methods of fluorescent biomolecular visualization, which required small molecule chemical administration. CRISPR is genetically programmable via nucleic acids, to target a specific genomic locus, making it easier to use than earlier methods of genome editing that required protein engineering. Finally, each has a clear mechanism of action, being implemented by a single well-understood protein, from a bioorthogonal species quite different from mammalian cells commonly studied in biomedicine. In our modern era of genomic search, directed evolution, and AI-guided molecular design, perhaps the ‘generalized Crick criteria’ could be coded up, or even automated, to accelerate the search for new molecular tools to advance biology and medicine.
Fourth, the technology should control neural ‘firing’ specifically, with a clear mechanism of action, so that there were no concerns about whether an unknown but required intermediary protein or other gene product was present or absent in a given cell type, or whether such an intermediary protein or other gene product could cause side effects, by coupling to unexpected physiological effectors in a given cell type.
Would such a technology be possible to create, and would it truly be useful in everyday neuroscience?
As a student at Stanford in spring 2000, one of us (Boyden) met another student, Karl Deisseroth, and we started brainstorming about how one might control neural activity in specific cell types by equipping targeted neurons with genetically encoded molecules that would transduce different forms of energy, such as magnetic fields, into electrical signals.
Reading old papers (Oesterhelt and Stoeckenius, Reference Oesterhelt and Stoeckenius1971; Schobert and Lanyi, Reference Schobert and Lanyi1982; Hildebrandt et al., Reference Hildebrandt, Fendler, Heberle, Hoffmann, Bamberg and Büldt1993; Hoffmann et al., Reference Hoffmann, Hildebrandt, Heberle and Büldt1994; Nagel et al., Reference Nagel, Möckel, Büldt and Bamberg1995; Okuno et al., Reference Okuno, Asaumi and Muneyuki1999), I became fascinated by the possibility of expressing microbial rhodopsins in neurons to make them sensitive to light and started requesting clones of such genes from colleagues (for a behind-the-scenes look at these early days, please see Boyden (Reference Boyden2011). I started with the light-driven chloride pump N. pharaonis halorhodopsin, because of a curious article suggesting that this protein might pump chloride well at modest salt concentrations (similar to those found in the brain, I noted at the time), in contrast to other microbial rhodopsins that worked best at high salt concentrations (perhaps because at the time, the best-studied microbial rhodopsins had been isolated from ‘halophilic’ archaea that live in high salinity environments) (Okuno et al., Reference Okuno, Asaumi and Muneyuki1999). That May, I emailed a request for this gene to Janos Lanyi, an opsin pioneer, who forwarded my request to his colleague Richard Needleman, who kindly sent the gene over. I had already headed out for the summer to a neuroscience course at the Marine Biology Laboratory in Woods Hole, Massachusetts. I asked Richard to send the gene to Karl. After returning to Stanford in fall 2000, I found myself rapidly caught up in learning lots of new skills in order to perform my PhD research on motor learning in the cerebellum, conducted in the labs of Jennifer Raymond and Dick Tsien, my PhD coadvisors, and I left the opsin project on the back burner for a while.
In the fall of 2003 and early 2004, Karl, then doing postdoctoral work in Rob Malenka’s lab, and I, mid-way through my PhD in Jennifer’s and Dick’s labs, started discussing genetically targeted neural control again. I had noticed a paper by Nagel et al. (Reference Nagel, Szellas, Huhn, Kateriya, Adeishvili, Berthold, Ollig, Hegemann and Bamberg2003), describing a light-activated cation channel, channelrhodopsin-2, and showing that this protein could be functionally expressed in oocytes or cultured HEK cells. I emailed Karl to propose that we reach out to Georg to see if they would be willing to share the gene. Georg kindly shared the gene, and we expressed it in cultured mammalian neurons.
There were many ways this experiment could have gone wrong – perhaps the protein could have been toxic to neurons, or perhaps the protein would not have functioned in neurons (perhaps it misfolded, or otherwise was compromised), or perhaps the effects would be too weak to be biologically meaningful. Or perhaps the protein would require the all-trans-retinal chemical cofactor to be supplemented, making usage too complex for everyday use in the alert mammalian brain. But amazingly, and serendipitously, it worked on the very first try!
On August 4, 2004, around 1 o’clock in the morning, working in Dick Tsien’s lab, I took a channelrhodopsin-2-expressing cultured mammalian neuron, began to electrophysiologically record it, and shined blue light on it – and to my amazement, it fired action potentials rapidly, precisely, and immediately. That night’s experiments confirmed that channelrhodopsin-2 was well-expressed, and functional, in neurons. The protein was well-tolerated enough by neurons, that it could be expressed at high levels, enough to mediate strong depolarizations. Brief pulses of blue light resulted in single, precisely timed action potentials in neurons, and trains of such pulses could result in precisely timed trains of action potentials. Repeatedly stimulating a neuron did not seem to cause a reduction in the opsin’s performance, suggesting that such optical control of neurons could be sustained over behaviorally relevant time periods. Serendipity had struck!
Follow-on experiments in the months to come, many performed in the Tsien lab, reinforced the excitement of that first night’s experimentation: the molecule was safe, functional, and effective. In August 2005, Karl and I published a paper reporting that the light-gated cation channel channelrhodopsin-2 from C. reinhardtii, expressed in cultured mammalian neurons, met all four of the criteria that Crick laid out (Boyden et al., Reference Boyden, Zhang, Bamberg, Nagel and Deisseroth2005). First, the small gene encoding for this protein could be genetically expressed in targeted neurons, using standard gene delivery and gene expression strategies common in biology. Second, the protein, expressed in neurons, was fast enough to mediate millisecond-timescale action potentials, in response to pulses of blue light. Third, the protein was easy to use in neurons, for example responding to blue light from a standard GFP excitation filter on a conventional microscope.
Most serendipitously, perhaps, the obligate chemical cofactor all-trans-retinal did not need to be supplemented to mammalian neurons – for whatever reason, mammalian neurons had sufficient background levels of all-trans-retinal to enable the function of microbial rhodopsins, which greatly simplified experiments. This serendipity ( Box 2) was reminiscent of how GFP spread quickly in biology in part because it required no chemical cofactors to be supplemented for its function (in contrast to some other biologically targeted fluorescent labeling schemes of the time).
Box 2. Optogenetics and the need for chemical supplementation.
As noted earlier ( Box 1), part of the utility of GFP arose from its ease of use – no chemicals needed to be supplemented for its everyday biological use in cells and organisms. In our original 2005 paper on the first use of microbial rhodopsins to mediate optical activation of neurons, we noted in the Discussion and Methods sections that ‘no all-trans retinal was added either to the culture medium or recording solution for any of the experiments described here’, expressing surprise that mammalian neurons seemed to do just fine with opsin functionality, even without adding all-trans-retinal. This turned out to be important for the ease of use of optogenetics in everyday neuroscience: if optogenetics required gene delivery to the living mammalian brain, implantation of an optical fiber (say, one that could be connected to an external light source) to target the region of interest with pulses of light, and then, on top of all that, infusion (either continuously or at time of experiment) of all-trans-retinal into the target region of the brain, the experiments would have been much more complicated than if only gene delivery (quite routine in neuroscience) and optical fiber implantation (analogous to ordinary electrode implantation) were required. In what might be regarded as a close call, although mammalian neurons did not require supplementation with all-trans-retinal for microbial rhodopsins to function, optogenetics does not work in the worm C. elegans or the fruit fly D. melanogaster without all-trans-retinal supplementation; fortunately, for these small animals, all-trans-retinal can be easily supplemented in sufficient quantities by adding it to the environment or to the food (Nagel et al., Reference Nagel, Brauner, Liewald, Adeishvili, Bamberg and Gottschalk2005; Schroll et al., Reference Schroll, Riemensperger, Bucher, Ehmer, Völler, Erbguth, Gerber, Hendel, Nagel, Buchner and Fiala2006).
Finally, since the protein directly coupled light to ion flux, without the need for another intermediary protein to achieve this coupling in neurons, there were no concerns about such intermediary proteins being potentially lacking in some neuron types, or about such intermediary proteins potentially causing side effects through coupling to unexpected downstream effectors. Thus, channelrhodopsin-2 fully enabled half of Crick’s proposed goal, specifically in the domain of neural activation. Several other papers using channelrhodopsin-2 in mouse brain slices, chick spinal cord, the worm C. elegans, and the mouse retina, came out in the months following, confirming these four properties of channelrhodopsin-2 in different contexts (Li et al., Reference Li, Gutierrez, Hanson, Han, Mark, Chiel, Hegemann, Landmesser and Herlitze2005; Nagel et al., Reference Nagel, Brauner, Liewald, Adeishvili, Bamberg and Gottschalk2005; Bi et al., Reference Bi, Cui, Ma, Olshevskaya, Pu, Dizhoor and Pan2006; Ishizuka et al., Reference Ishizuka, Kakuda, Araki and Yawo2006).
In 2007, after I started my group at MIT, Xue Han and I showed that the light-driven chloride pump N. pharaonis halorhodopsin – the very first microbial opsin clone I requested from colleagues, back in the spring of 2000 – possessed these four properties in the domain of neural silencing (Han and Boyden, Reference Han and Boyden2007). Our paper was followed shortly after, by a paper on the same molecule, from the Deisseroth lab (Zhang et al., Reference Zhang, Wang, Brauner, Liewald, Kay, Watzke, Wood, Bamberg, Nagel, Gottschalk and Deisseroth2007). The silencing was not very strong, however, perhaps because the halorhodopsin was not functionally expressed at high enough levels in mammalian neurons. In 2010, my group at MIT showed that a light-driven proton pump from H. sodomense, archaerhodopsin-3, could mediate much more powerful neural silencing, with ~100% reduction of neural firing in cortical neurons of awake behaving mice in response to pulses of light (Chow et al., Reference Chow, Han, Dobry, Qian, Chuong, Li, Henninger, Belfort, Lin, Monahan and Boyden2010), followed shortly after by a paper from the Deisseroth lab showing that the photocurrents of the N. pharaonis halorhodopsin could be improved by adding trafficking signals that boosted neural functional expression (Gradinaru et al., Reference Gradinaru, Zhang, Ramakrishnan, Mattis, Prakash, Diester, Goshen, Thompson and Deisseroth2010).
These molecules, thus, enabled the other half of Crick’s proposed goal, specifically in the domain of neural silencing. They remain popular to this day. We, and many others, have continued to discover new molecules that are more optimal for specialized purposes (discussed at length in the final parts of this review) – as just a few examples, enabling very fast neural control (Klapoetke et al., Reference Klapoetke, Murata, Kim, Pulver, Birdsey-Benson, Cho, Morimoto, Chuong, Carpenter, Tian, Wang, Xie, Yan, Zhang, Chow, Surek, Melkonian, Jayaraman, Constantine-Paton, Wong and Boyden2014), enabling less invasive neural control (Han et al., Reference Han, Chow, Zhou, Klapoetke, Chuong, Rajimehr, Yang, Baratta, Winkle, Desimone and Boyden2011; Chuong et al., Reference Chuong, Miri, Busskamp, Matthews, Acker, Sørensen, Young, Klapoetke, Henninger, Kodandaramaiah, Ogawa, Ramanlal, Bandler, Allen, Forest, Chow, Han, Lin, Tye, Roska, Cardin and Boyden2014; Klapoetke et al., Reference Klapoetke, Murata, Kim, Pulver, Birdsey-Benson, Cho, Morimoto, Chuong, Carpenter, Tian, Wang, Xie, Yan, Zhang, Chow, Surek, Melkonian, Jayaraman, Constantine-Paton, Wong and Boyden2014), enabling multiplexed neural control (Klapoetke et al., Reference Klapoetke, Murata, Kim, Pulver, Birdsey-Benson, Cho, Morimoto, Chuong, Carpenter, Tian, Wang, Xie, Yan, Zhang, Chow, Surek, Melkonian, Jayaraman, Constantine-Paton, Wong and Boyden2014), enabling ion-selective neural control (Cho et al., Reference Cho, Yong, Yang, Chen, Chuong, Klapoetke and Boyden2019), and enabling very spatially precise neural control (Shemesh et al., Reference Shemesh, Tanese, Zampini, Linghu, Piatkevich, Ronzitti, Papagiakoumou, Boyden and Emiliani2017), amongst others – which are also helping neuroscientists tackle a great many specialized problems.
In summary, the class of microbial rhodopsins, with little or no modification from their natural state, was able to address a key need in neuroscience, enabling the fast, easy-to-use, and reliable activation and silencing of electrical activity in specific neuron types, in response to light. This was largely due to serendipity: the molecules might not have been fast or strong enough, when embedded in the neuronal milieu, to mediate neural firing, or they may have proven toxic in delicate mammalian neurons, or they may have required chemical supplementation of all-trans-retinal to function in mammalian neurons, greatly complicating experimentation.
In the years since, optogenetic tools have been used in practically every part of neuroscience to study how the activities of specific cell types contribute to behaviors, pathological states, or potential therapeutic processes. Because the tools are easy to express in targeted neurons, using standard gene delivery and transgenesis strategies, they are widely used in the major model organisms utilized in neuroscience, including mice, rats, nonhuman primates, flies, fish, and worms. We have distributed these tools as freely as possible to the neuroscience community, for example, working with DNA-repository services like Addgene to distribute plasmids, and viral vector cores at many different institutions to distribute viruses. They have been used by perhaps thousands of researchers in animals to probe literally hundreds of topics related to normal and pathological brain states and processes. It is probably impossible to list all of the papers that utilize optogenetics and still maintain a cohesive review, especially one focused on the biophysics of the rhodopsins in their neuroscience roles, but in the next section we try to give a flavor for the kinds of results people have obtained, using optogenetic tools in neuroscience, before moving onto discussion of the biophysical details of optogenetics and how these properties helped these tools stand out in neuroscience utility.
Application of rhodopsins in neuroscience
In this section, we give examples of the kind of results scientists have obtained, first in basic science studies of how neurons work together in circuits to generate behavior in a variety of model organisms, and then in studies to probe the nature of brain diseases and to think about new strategies to treat them. Although the promise of optogenetics has paid off hugely in the understanding of the brain, revealing the causal substrates of a great many behaviors and diseases, and pointing in many cases toward potential new treatment strategies, a second major potential impact – direct application of optogenetics in humans, as a therapeutic – is starting to be substantiated by data from human patients with blindness, and may represent a second payoff of optogenetics; we discuss this new direction briefly at the end of this section.
Optogenetic tools have been used in mammals including mice and rats to reveal neural populations and activity patterns that drive parental behaviors (Kohl et al., Reference Kohl, Babayan, Rubinstein, Autry, Marin-Rodriguez, Kapoor, Miyamishi, Zweifel, Luo, Uchida and Dulac2018), that enhance spatial object recognition (Kempadoo et al., Reference Kempadoo, Mosharov, Choi, Sulzer and Kandel2016), that drive attacks upon intruders (Lin et al., Reference Lin, Boyle, Dollar, Hyosang Lee, Perona and Anderson2011), that control the timing of breathing (Sherman et al., Reference Sherman, Worrell, Cui and Feldman2015), that are needed for social memory formation (Oliva et al., Reference Oliva, Fernández-Ruiz, Leroy and Siegelbaum2020), that regulate the formation of social-spatial associations (Murugan et al., Reference Murugan, Jang, Park, Miller, Cox, Taliaferro, Parker, Bhave, Hur, Liang, Nectow, Pillow and Witten2017), that improve visual perception (Lee et al., Reference Lee, Kwan, Zhang, Phoumthipphavong, Flannery, Masmanidis, Taniguchi, Huang, Zhang, Boyden, Deisseroth and Dan2012), that boost wakefulness (Cho et al., Reference Cho, Treweek, Robinson, Xiao, Bremner, Greenbaum and Gradinaru2017), that control locomotor-like bursting in spinal cord central pattern generators (Hagglund et al., Reference Hagglund, Dougherty, Borgius, Itohara, Iwasato and Kiehn2013), that control the duration and physiological properties of sleep episodes (Jego et al., Reference Jego, Glasgow, Herrera, Ekstrand, Reed, Boyce, Friedman, Burdakov and Adamantidis2013), that are necessary for formation of long-term memories (Kitamura et al., Reference Kitamura, Ogawa, Roy, Okuyama, Morrissey, Smith, Redondo and Tonegawa2017), that encode the laterality of sensory inputs (Ketzef et al., Reference Ketzef, Spigolon, Johansson, Bonito-Oliva, Fisone and Silberberg2017), that contribute to goal-directed attentional processing (Kim et al., Reference Kim, Ährlund-Richter, Wang, Deisseroth and Carlén2016), that play a causal role in face gender discrimination (Afraz et al., Reference Afraz, Boyden and DiCarlo2015), that are necessary for driving water consumption in conditions of thirst (Zimmerman et al., Reference Zimmerman, Lin, Leib, Guo, Huey, Daly, Chen and Knight2016), that recapitulate innate responses to odors (Root et al., Reference Root, Denny, Hen and Axel2014), that control food intake in conditions of hunger (Nectow et al., Reference Nectow, Schneeberger, Zhang, Field, Renier, Azevedo, Patel, Liang, Mitra, Tessier-Lavigne, Han and Friedman2017), that modulate specific aspects of movement (Gritton et al., Reference Gritton, Howe, Romano, DiFeliceantonio, Kramer, Saligrama, Bucklin, Zemel and Han2019), that control memory-guided eye movements (Acker et al., Reference Acker, Pino, Boyden and Desimone2016), that induce aversion or preference to a place (Kim et al., Reference Kim, Lee, Fang, Shin, Park, Hashikawa, Bhat, Kim, Sohn, Lin and Suh2019), that promote conditioned reward-seeking behavior (Otis et al., Reference Otis, Vijay, Matan, Voets, Mohorn, Kosyk, McHenry, Robinson, Resendez, Rossi and Stuber2017), that regulate paternal behavior (Stagkourakis et al., Reference Stagkourakis, Smiley, Williams, Kakadellis, Ziegler, Bakker, Rosemary, Harkany, Grattan and Broberger2020), and that provide signals to the hippocampus to help neurons encode for places (Zhang et al., Reference Zhang, Ye, Miao, Tsao, Cerniauskas, Ledergerber, Moser and Moser2013) – amongst countless other results.
In important small model organisms for neuroscience, optogenetics has proven very useful in defining neural populations and activity patterns that contribute to neural computations and behaviors.
In fruit flies, optogenetics has been used to reveal neural populations and activity patterns that control acquired feeding preferences (Musso et al., Reference Musso, Junca, Jelen, Feldman-Kiss, Zhang, Rachel and Gordon2019), that control chemotactic navigational decision-making (Hernandez-Nunez et al., Reference Hernandez-Nunez, Belina, Klein, Si, Claus, Carlson and Aravinthan2015), that drive or inhibit courtship (Seeholzer et al., Reference Seeholzer, Seppo, Stern and Ruta2018), that promote sleep and suppress locomotor activity (Guo et al., Reference Guo, Yu, Jung, Abruzzi, Luo, Griffith and Rosbash2016), that drive a long-lasting internal state in the female brain that regulates a diverse set of behaviors (Deutsch et al., Reference Deutsch, Pacheco, Encarnacion-Rivera, Pereira, Fathy, Clemens, Girardin, Calhoun, Ireland, Burke, Dorkenwald, McKellar, Macrina, Lu, Lee, Kemnitz, Ih, Castro, Halageri, Jordan, Silversmith, Wu, Seung and Murthy2020), that process touch signals in a set of parallel comparisons (Tuthill and Wilson, Reference Tuthill and Wilson2016), that control context-appropriate walking programs (Bidaye et al., Reference Bidaye, Laturney, Chang, Liu, Bockemühl, Büschges and Scott2020), that result in a diversity of complex and novel behavioral sequences (Vogelstein et al., Reference Vogelstein, Park, Ohyama, Kerr, Truman, Priebe and Zlatic2014), and that represent the heading direction of a fly-through ring attractor dynamics (Kim et al. Reference Kim, Bae, Capece, Nedelkovska, de Rubio, Smrcka, Jun, Jung, Park, Kim and Kim2017), amongst many other discoveries.
In the larval zebrafish, optogenetics revealed neural populations and activity patterns that controlled saccadic eye movements (Schoonheim et al., Reference Schoonheim, Arrenberg, del Bene and Baier2010), that increase sleep (Oikonomou et al., Reference Oikonomou, Altermatt, Zhang, Coughlin, Montz, Gradinaru and Prober2019), that control swim turn direction (Dunn et al., Reference Dunn, Mu, Narayan, Randlett, Naumann, Yang, Schier, Freeman, Engert and Ahrens2016), that provide sensory feedback to spinal circuits during fast locomotion (Knafo et al., Reference Knafo, Fidelin, Prendergast, Tseng, Parrin, Dickey, Böhm, Figueiredo, Thouvenin, Pascal-Moussellard and Wyart2017), that produce a coordinated swimming pattern (Ljunggren et al., Reference Ljunggren, Haupt, Ausborn, Ampatzis and el Manira2014), that stop ongoing swimming (Kimura et al., Reference Kimura, Satou, Fujioka, Shoji, Umeda, Ishizuka, Yawo and Higashijima2013), and that contribute to movement in response to noxious stimuli (Wee et al., Reference Wee, Nikitchenko, Wang, Luks-Morgan, Song, Gagnon, Randlett, Bianco, AMB, Glushenkova, Barrios, Schier, Kunes, Engert and Douglass2019), amongst other discoveries.
In the worm C. elegans, optogenetics has been used to pinpoint neurons involved with generating locomotor rhythms (Fouad et al., Reference Fouad, Teng, Mark, Liu, Alvarez-Illera, Ji, Du, Bhirgoo, Cornblath, Guan and Fang-Yen2018), and to explore how a single neuron can regulate multiple behavioral outputs (Li et al., Reference Li, Liu, Zheng and Xu2014), how specific neurons mediate the switching of behavioral state in response to oxygen concentrations reflective of surface exposure (Laurent et al., Reference Laurent, Soltesz, Nelson, Chen, Arellano-Carbajal, Levy and de Bono2015), how interneurons integrate multiple kinds of olfactory input toward a representation of valence (Dobosiewicz et al., Reference Dobosiewicz, Liu and Bargmann2019), how a single neuron encodes a memory of a chemotactic set point (Luo et al., Reference Luo, Wen, Ren, Hendricks, Gershow, Qin, Greenwood, Soucy, Klein, Smith-Parker, Calvo, Colón-Ramos, Samuel and Zhang2014), how synaptic energy demand regulates the clustering of a glycolytic protein (Jang et al., Reference Jang, Nelson, Bend, Rodríguez-Laureano, Tueros, Cartagenova, Underwood, Jorgensen and Colón-Ramos2016), how specific neurons contribute oscillatory activity to control backward locomotion (Gao et al., Reference Gao, Guan, Fouad, Meng, Kawano, Huang, Li, Alcaire, Hung, Lu, Qi, Jin, Alkema, Fang-Yen and Zhen2018), and how specific interneurons control the locomotory programs for chemotaxis (Kocabas et al., Reference Kocabas, Shen, Guo and Ramanathan2012) – again, amongst a large number of studies from all over the world.
Beyond the most commonly used model organisms in neuroscience, optogenetics has also been applied to the study of neural circuits and behaviors in other species important in neuroscience, including nonhuman primates (Han et al., Reference Han, Qian, Bernstein, Zhou, Franzesi, Stern, Bronson, Graybiel, Desimone and Boyden2009).
Optogenetics has also been used to study diseases in animal models of brain disorders, pinpointing cell types and neural circuits that could serve as therapeutic targets for treating brain diseases, and even revealing neural activity patterns that could, when induced by brain stimulation technology, potentially serve therapeutic roles. Optogenetic control of neurons has revealed, in animal species and models relevant to human diseases and conditions, neural populations and activity patterns that clean up multiple molecular pathologies associated with Alzheimer’s disease (Iaccarino et al., Reference Iaccarino, Singer, Martorell, Rudenko, Gao, Gillingham, Mathys, Seo, Kritskiy, Abdurrob, Adaikkan, Canter, Rueda, Brown, Boyden and Tsai2016), that wake the brain up from anesthesia (Taylor et al., Reference Taylor, Van Dort, Kenny, Pei, Guidera, Vlasov, Lee, Boyden, Brown and Solt2016), that relieve anxiety-like states in stressed mice (Kumar et al., Reference Kumar, Black, Hultman, Szabo, Demaio, Du, Katz, Feng, Covington and Dzirasa2013), that control the acquisition of learned fear (Wolff et al., Reference Wolff, Jan Gründemann, Tovote, Krabbe, Jacobson, Müller, Herry, Ehrlich, Friedrich, Letzkus and Lüthi2014) or the encoding of contextual fear memories (Kheirbek et al., Reference Kheirbek, Drew, Burghardt, Costantini, Tannenholz, Ahmari, Zeng, Fenton and Henl2013), that control the generalization of fear memories (Xu and Sudhof, Reference Xu and Sudhof2013), that promote compulsive seeking of sugar (Nieh et al., Reference Nieh, Matthews, Allsop, Presbrey, Leppla, Wichmann, Neve, Wildes and Tye2015), that promote spinal cord repair after injury (Llorens-Bobadilla et al., Reference Llorens-Bobadilla, Chell, le Merre, Wu, Zamboni, Bergenstråhle, Stenudd, Sopova, Lundeberg, Shupliakov, Carlén and Frisén2020), that restore respiratory diaphragm motor activity after spinal cord injury (Alilain et al., Reference Alilain, Li, Horn, Dhingra, Dick, Herlitze and Silver2008), that drive depression-like behaviors (Yang et al., Reference Yang, Cui, Sang, Dong, Ni, Ma and Hu2018), that participate in or promote post-stroke motor recovery (Cheng et al., Reference Cheng, Wang, Woodson, Wang, Sun, Lee, Arac, Fenno, Deisseroth and Steinberg2014; Wahl et al., Reference Wahl, Büchler, Brändli, Brattoli, Musall, Kasper, Ineichen, Helmchen, Ommer and Schwab2017), that are dysregulated in states of obesity (Pirzgalska et al., Reference Pirzgalska, Seixas, Seidman, Link, Sánchez, Mahú, Mendes, Gres, Kubasova, Morris, Arús, Larabee, Vasques, Tortosa, Sousa, Anandan, Tranfield, Hahn, Iannacone, Spann, Glass and Domingos2017; Reed et al., Reference Reed, Yim, Wimmer, Kim, Ryu, Welch, Andina, King, Waisman, Halassa, Huh and Choi2019; Beutler et al., Reference Beutler, Corpuz, Ahn, Kosar, Song, Chen and Knight2020), that normalize motor behavior in Huntington’s model mice (Fernández-García et al., Reference Fernández-García, Conde-Berriozabal, García-García, Gort-Paniello, Bernal-Casas, Barriga, López-Gil, Muñoz-Moreno, Soria, Campa, Artigas, Rodríguez, Alberch and Masana2020), that cause long-lasting motor recovery in dopamine-depleted mice (Mastro et al., Reference Mastro, Zitelli, Willard, Leblanc, Kravitz and Gittis2017), that control cocaine-seeking behavior relevant to addiction (Martín-García et al., Reference Martín-García, Courtin, Renault, Fiancette, Wurtz, Simonnet, Levet, Herry and Deroche-Gamonet2014), that disrupt the role of sleep in consolidating memories (Swift et al., Reference Swift, Gross, Frazer, Bauer, Kyle, Vazey, Aston-Jones, Pickering, Sara and Poe2018), that inhibit epileptic bursting in hippocampal and cortical brain circuits (Tonnesen et al., Reference Tonnesen, Sorensen, Deisseroth, Lundberg and Kokaia2009), that stop seizures in vivo (Krook-Magnuson et al., Reference Krook-Magnuson, Armstrong, Oijala and Soltesz2013), that halt seizures that result from stroke (Paz et al., Reference Paz, Davidson, Frechette, Delord, Parada, Peng, Deisseroth and Huguenard2013), that ameliorate Parkinsonian motor symptoms (Yu et al., Reference Yu, Cassar, Sambangi and Grill2020), that contribute to stem-cell derived reduction of Parkinson’s symptoms (Steinbeck et al., Reference Steinbeck, Choi, Mrejeru, Ganat, Deisseroth, Sulzer, Mosharov and Studer2015), and that overcome developmental limitations on social learning (Nardou et al., Reference Nardou, Lewis, Rothhaas, Xu, Yang, Boyden and Dölen2019) – again, out of a great many clinically informative results from a large number of groups.
The widespread usage of optogenetics in awake-behaving animals has been greatly facilitated by the utility of ordinary laser, LED, fiber optic, and microscopy technology, to deliver light to the brain, effectively, easily, and safely. Many of the mouse studies above, for example, involved implanting an optic fiber into the brain, with one end aimed at a brain region of interest. The brain region of interest will typically have had one cell type of interest made sensitive to light through the expression of an appropriate light-activated pump or channel in the cell type of interest, using standard gene delivery mechanisms (e.g., an AAV virus, containing the gene encoding for a given opsin, perhaps under regulatory sequences to help a specific cell type express the gene selectively, could be stereotactically injected into the region of interest). At the time of a behavioral experiment, the other end of the optic fiber, which emerges from the brain, would be connected to an external LED or laser of the appropriate color, which would then be pulsed by a computer, to drive the neural code according to some experimental goal. For small animals like worms, flies, and fish, they are often simply placed under a standard microscope, which then delivers light of appropriate color and timing, to the brain or body. Transgenic methods will have been used to enable specific cell types, in the brain or body, to express the rhodopsins.
As optics hardware improves over time – for example, multiphoton, digital micromirror device, and holographic light sculpting hardware, have been making their way more and more into neuroscience in recent years, to facilitate neural imaging – such devices are being adapted for making optogenetic control more and more spatially precise, as well. Reviewing the optical hardware of optogenetics is beyond the scope of this review, which is focused on the chemistry and biophysics of the molecules and how those properties yielded impact on neuroscience. Although our focus in the aforementioned examples has been on the application of optogenetics in the intact brain, often in behaving animals, we note that countless studies in vitro, including studies of mechanisms of neural communication, intraneuronal computation, neural plasticity, circuit organization, and circuit dynamics, performed using acute brain slices and other in vitro preparations such as cultured primary neurons, as well as in many nonneuronal systems comprising excitable cells such as heart and musculature, have been enabled by optogenetics as well.
Optogenetics has had an enormous impact on the study of the brain, pointing to cell types, neural circuits, and neural codes that causally contribute to a diversity of behaviors, disease states, and potential therapeutic states. In this regard, optogenetic usage is proven and mainstream, and is now routinely used in everyday neuroscience to probe the cellular and circuit mediators of normal and abnormal neural processes. In the last few years, however, a second frontier has begun to gain more attention – the potential for directly using optogenetics in human patients, to treat diseases or restore function. For optogenetics to be used in a human patient, since it would require both a gene therapy to introduce the gene into specific cells in the body, as well as a hardware device for controlled light delivery to target cells, there would need to be a rationale for a specific cell type or neural circuit target to be selected to express the optogenetic molecule; there would need to be optical hardware to deliver light of the appropriate color and power to the region of interest, precisely and safely; and there would need to be preclinical data as well as clinical trials to support both the safety of the molecule in the body (since they evolved in species very different from humans, a lack of toxicity of the gene product, and a lack of immune response against it, ideally over timescales relevant to human disease treatment, would have to be confirmed) as well as the efficacy of the neural modulation in ameliorating the condition or restoring function.
In summer 2021, these goals converged for the first time in a human patient (Sahel et al., Reference Sahel, Boulanger-Scemama, Pagot, Arleo, Galluppi, Martel, Esposti, Delaux, de Saint Aubert, de Montleau, Gutman, Audo, Duebel, Picaud, Dalkara, Blouin, Taiel and Roska2021), with the first case study being reported of a patient suffering from retinitis pigmentosa, a disease that causes photoreceptor loss and resulting blindness, achieving a partial restoration of functional vision after AAV-mediated delivery of the gene encoding for the light-driven cation channel ChrimsonR (Klapoetke et al., Reference Klapoetke, Murata, Kim, Pulver, Birdsey-Benson, Cho, Morimoto, Chuong, Carpenter, Tian, Wang, Xie, Yan, Zhang, Chow, Surek, Melkonian, Jayaraman, Constantine-Paton, Wong and Boyden2014) into the eye, targeting normally light-insensitive retinal ganglion cells, to make them light-sensitive. In this way, the retina could convert light into neural signals for relay on to the brain, even though the natural photoreceptors were gone. The patient wore goggles that captured images of the world and projected processed images in the form of patterned light pulses of appropriate color and power, to the retina. In this patient, there were no adverse events reported. Tantalizingly, there was significant restoration of functional vision, including the ability to perceive, reach for, and touch objects, to the point of being able to perform some daily visual activities – perceiving crosswalks and doors on the street and in hallways respectively, and detecting household objects like plates and phones. Perception persisted over the duration of the study (over 1.5 years of testing). Future studies will be needed, in a larger cohort of patients, both in the context of this disease and in any diseases to be explored in the future, to fully understand the potential of optogenetics in direct treatment of human diseases and in restoration of function.
The landscape of pioneering neural control technologies
In the years before 2005, when the first use of microbial rhodopsins to mediate optical control of neural activity was published, many pioneering scientists and engineers worked on innovative strategies to enable neural control that was more precise than classical pharmacology and electrical stimulation. Each of these techniques met a subset of the four criteria mentioned above, so although none of these techniques spread throughout neuroscience at the time, they validated key aspects of the concept of precision neuron control. In this section, we briefly review the landscape of precision neural control in 2005 and before, going over different classes of technology and what aspects of neural control they enabled. Although many of these classes of tool have improved post-2005 and some are now in widespread use in neuroscience, reviewing these post-2005 improvements and inventions is beyond the scope of this review, which is focused on optogenetics and the biophysics thereof.
One class of methods involved the direct optical stimulation of neurons. Such techniques could be very fast, because they use light as the trigger, but given their reliance on endogenous, sometimes unclear, mechanisms of action, it could be hard to judge how well a given technology could be targeted to different cell types, whether it would be generally easy to use, and whether unknown intermediaries were required that may not be universally available across different cell types, or that could engage pathways that cause side effects. Hints of the possibility of using light to directly control cellular excitability go back over a century; for example, one paper in 1891 reported the excitation of muscle fibers using light (Arsonval, Reference Arsonval1891). Following previous biophysical observations (Chalazonitis, Reference Chalazonitis1964), it was shown that shining visible laser light on neurons of Aplysia could be used to trigger neural activity with second-timescale latency (Fork, Reference Fork1971), with unclear mechanism of action (Allègre et al., Reference Allègre, Avrillier and Albe-Fessard1994). Another report showed that two-photon excitation could be used to activate neurons directly in mouse cortical brain slices (Hirase et al., Reference Hirase, Nikolenko, Goldberg and Yuste2002), with millisecond precision, although again the mechanism of action was unclear; one possibility the authors mentioned was the laser-induced formation of microholes in the membrane. Infrared light was also shown to be capable of directly exciting peripheral nerves in vivo in frogs and rats, potentially through a thermal effect (Wells et al., Reference Wells, Kao, Mariappan, Jeffrey Albea, Konrad and Mahadevan-Jansen2005).
A second class of methods used small-molecule chemicals to help mediate the conversion of light into a neural activating stimulus. Such techniques could again be very fast but could not be targeted to a specific neuron type, and the requirement for exogenous chemical delivery would require such delivery to occur in the living brain for behavioral use. Optical activation of neurons using light to uncage the neurotransmitter glutamate at sites in rat cortical and hippocampal brain slices (Callaway and Katz, Reference Callaway and Katz1993) resulted in millisecond-timescale neural activation of nearby neurons, with a clear mechanism of action since it simulated pulsatile transmitter presence. Another study showed that staining neurons from leeches, frogs, and other species with a specific small molecule dye resulted in laser-elicited action potentials within milliseconds (Farber and Grinvald, Reference Farber and Grinvald1983), with an unclear mechanism of action, although one possibility the authors mentioned is the transient formation of membrane channels.
A third class of methods used the genetic expression of an ion channel gene, or ion channel modulating gene, to perturb electrical activity in targeted cells. Such a strategy would be limited to a temporal precision associated with the rate of gene expression, but would be easy to use, requiring nothing beyond gene delivery to operate, and would have a clear mechanism of physiological action. Expressing natural or modified potassium channels that hyperpolarize neurons, using standard gene delivery, transgenesis, and/or inducible gene expression strategies, in mammalian neurons and other excitable cells in culture and in vivo, and in Aplysia, Xenopus, C. elegans, and Drosophila neurons and other excitable cells, enabled in many cases the electrical quieting or silencing of these cells (Kaang et al., Reference Kaang, Pfaffinger, Seth, Kandel and Furukawa1992; Jones and Ribera, Reference Jones and Ribera1994; Ehrengruber et al., Reference Ehrengruber, Doupnik, Xu, Garvey, Jasek, Lester and Davidson1997; Johns et al., Reference Johns, Marx, Mains, O’Rourke and Marbán1999; Peckol et al., Reference Peckol, Zallen, Yarrow and Bargmann1999; Sutherland et al., Reference Sutherland, Williams, Abedi, Overbeek, Pfaffinger and Noebels1999; Nadeau et al., Reference Nadeau, McKinney, Anderson and Lester2000; Baines et al., Reference Baines, Uhler, Thompson, Sweeney and Bate2001; Falk et al., Reference Falk, Kilani, Yool and Sherman2001; Paradis et al., Reference Paradis, Sweeney and Davis2001; White et al., Reference White, Osterwalder, Yoon, Joiner, Whim, Kaczmarek and Keshishian2001; Burrone et al., Reference Burrone, O’Byrne and Murthy2002; Nitabach et al., Reference Nitabach, Blau and Holmes2002; Yu et al., Reference Yu, Power, Barnea, O’Donnell, Brown, Osborne, Axel and Gogos2004) with a time precision of hours to days and no need for chemical supplementation, although some of these studies noted that long-term expression of such channels could cause various side effects and toxicities, perhaps as a consequence of extremely long duration hyperpolarization. Expressing an appropriately mutated glutamate receptor in specific C. elegans neurons caused them to be activated, and for specific behaviors to be elicited (Zheng et al., Reference Zheng, Brockie, Mellem, Madsen and Maricq1999). Another study showed that tethering to the cell membrane ion channel-blocking toxins that blockade sodium channels, calcium channels, and other channels, could be achieved in a genetically encoded construct; in living zebrafish, such a strategy was used to block cholinergic receptors (Ibañez-Tallon et al., Reference Ibañez-Tallon, Wen, Miwa, Xing, Tekinay, Ono, Brehm and Heintz2004).
A fourth class of methods used a gene that encoded for an ion channel, which could then be actuated by a chemical (‘chemogenetics’). A related class of method used a gene that encoded for an ion channel that could be equipped with a chemical and then actuated using light (‘photopharmacogenetics’). The time precision of chemogenetics would be related to the adding or removing of the chemical; the time precision of photopharmacogenetics would be related to the timescale of the delivery of light. Cell type targetability would be facilitated by the genetic nature of the ion channel; delivery of a chemical must be achieved for use in the living brain. The mechanism of action would be as clear as the understanding of the nature of the ion channel biology and of the chemical ligand. One study virally delivered the C. elegans chloride channel GluCl to cultured rat hippocampal neurons and showed that the drug ivermectin could be used to silence their electrical activity (Slimko et al., Reference Slimko, McKinney, Anderson, Davidson and Lester2002); the time to achieve silencing was seconds. Another study showed that genetic delivery of a potassium channel engineered to bind a photoswitchable tethered pore blocker (building from earlier studies on using photoswitchable tethered ligands to activate ion channel proteins such as cholinergic receptors (Bartels et al., Reference Bartels, Wassermann and Erlanger1971; Lester et al., Reference Lester, Krouse, Nass, Wassermann and Erlanger1980)) to cultured hippocampal neurons, followed by the delivery of the photoswitchable tethered pore blocker, enabled these neurons to be activated by light within seconds (Banghart et al., Reference Banghart, Borges, Isacoff, Trauner and Kramer2004). In another study, expressing the capsaicin-activated cation channel TRPV1 in a specific neuron in C. elegans, and exposing the worm to capsaicin, caused behaviors consistent with the activation of the targeted neuron (Tobin et al., Reference Tobin, Madsen, Kahn-Kirby, Peckol, Moulder, Barstead, Maricq and Bargmann2002). In another study, investigators expressed ion channels that are gated by agonists not naturally found in the nervous system, such as the TRPV1 channel or the P2X2 channel, in cultured hippocampal neurons, and then found that adding the agonists capsaicin or ATP respectively, or optically uncaging caged capsaicin or ATP onto, these neurons resulted within seconds in neural activity (Zemelman et al., Reference Zemelman, Nesnas, Lee and Miesenböck2003); by expressing the P2X2 channel in specific Drosophila neurons and injecting caged ATP into the central nervous system, light illumination was able to reveal behaviors triggered by activation of those neurons (Lima and Miesenböck, Reference Lima and Miesenböck2005).
A fifth class of methods used a gene that encodes for a signaling cascade molecule (sometimes with accessory proteins to help it function), most often a G-protein coupled receptor (GPCR), that could couple to downstream physiological effectors (such as endogenous ion channels). The GPCR could then be actuated by a chemical, for example, a ligand that binds the receptor. Alternatively, the GPCR could be equipped with a chemical and then actuated by light. As with the previous class, the time precision would be related to the adding or removing of the chemical, or by the delivery of light; cell type targetability would be facilitated by the genetic nature of the signaling cascade; delivery of a chemical must be achieved for use in the living brain. The mechanism of action could depend on the nature of the cell type being targeted; for signaling cascades downstream of a GPCR, unknown but required intermediary proteins may be present or absent in a given cell type, or such intermediary proteins could cause side effects by coupling to other, unexpected physiological effectors. However, such intermediaries may also amplify the impact of a chemical or optical trigger on neural physiology, increasing the amplitude of an effect. In one study, expression of a modified human kappa opioid GPCR in the mouse heart enabled, upon administration of the drug spiradoline, reduction of heart rate within seconds (Redfern et al., Reference Redfern, Coward, Degtyarev, Lee, Kwa, Hennighausen, Bujard, Fishman and Conklin1999); this GPCR signals through Gi, which in the heart inhibits adenylyl cyclase and activates a membrane potassium channel. By expressing the Drosophila allatostatin receptor, which exhibits Gi/o signaling, along with G-protein-coupled inwardly rectifying potassium (GIRK) channel subunits that are regulated by Gi/o (required because at the age of the brain being studied, such GIRK channels are not yet expressed), in cultured ferret visual cortex brain slices, neurons could be silenced within minutes of adding the ligand allatostatin (Lechner et al., Reference Lechner, Lein and Callaway2002). Another approach involved equipping cells with the gene for a G-protein coupled rhodopsin and a retinal cofactor. In one such study, frog oocytes received the gene for bovine rhodopsin and then were incubated with 11-cis-retinal; illumination caused engagement of the G protein Gt, and caused photocurrents within seconds (Khorana et al., Reference Khorana, Knox, Nasi, Swanson and Thompson1988). In another study, investigators expressed G-protein coupled Drosophila rhodopsin, arrestin-2, and the Gqalpha subunit of the downstream G protein cascade, in cultured hippocampal neurons, and added an initial dose of all-trans-retinal beyond background levels to reconstitute the rhodopsin (Zemelman et al., Reference Zemelman, Lee, Ng and Miesenböck2002); this rhodopsin signaled to available downstream effectors, ultimately opening available cation channels in cells in which they are expressed. Upon illumination, neural activity began within hundreds of milliseconds to tens of seconds. Three studies published almost on the same day showed that expressing human melanopsin in cultured mammalian cells, supplemented with 9-cis or 11-cis retinaldehyde, resulted, upon illumination, in G-protein-mediated photocurrents within seconds (Melyan et al., Reference Melyan, Tarttelin, Bellingham, Lucas and Hankins2005; Panda et al., Reference Panda, Nayak, Campo, Walker, Hogenesch and Jegla2005; Qiu et al., Reference Qiu, Kumbalasiri, Carlson, Wong, Krishna, Provencio and Berson2005).
We have focused our discussion above on pioneering tools that manipulated electrical activity in targeted cells, before 2005. Of course, manipulations of many other biological functions that affect neural signaling, including alteration of synapses or synaptic transmission in targeted cells, as well as ways of lesioning or killing targeted cells, have played major roles in neuroscience, both before and after 2005, but are beyond the scope of this review. In addition, this review is not intending to comprehensively review nonoptogenetic technologies for controlling targeted neural electrical activity after 2005, since the goal was to outline the landscape at the time, in hopes of exploring what biophysical properties of microbial rhodopsins led to optogenetics taking off. Many nonoptogenetic toolsets for controlling targeted neural electrical activity, including novel toolsets (e.g., magnetogenetics, sonogenetics), as well as extensions of the aforementioned ones (e.g., chemogenetics), have exploded in utility and popularity since 2005, in their own right, both because of continued ingenious engineering and resulting improved performance, as well as availability of synergistic tools (e.g., viral gene delivery and the availability of viruses from core facilities has facilitated the deployment and use of a great many such genetically encoded tools throughout neuroscience).
The landscape of opsin discovery and application
We here review the microbial opsin discoveries that preceded the adaptation of microbial rhodopsins for mediating the optical control of neural electrical activity; the section following will review the biophysical properties of rhodopsins that conferred their utility for specific neuroscience experiments. Microbial rhodopsins were first reported in the early 1970s, with the discovery of bacteriorhodopsin, a protein in the halophilic archaeal species Halobacterium salinarum (formerly known as Halobacterium halobium) that was found to be a rhodopsin-like protein, a membrane protein that bound retinal and that exhibited particular compositional and spectral properties, and that pumped protons outwards across cellular membranes in response to light (Oesterhelt and Stoeckenius, Reference Oesterhelt and Stoeckenius1971, Reference Oesterhelt and Stoeckenius1973), helping store the energy of sunlight in a chemical gradient for downstream ATP production (Danon and Stoeckenius, Reference Danon and Stoeckenius1974).
Around a decade later, a second rhodopsin-like protein, a light-driven chloride pump, named halorhodopsin, was discovered in the same species of archaea, where it also contributes to bioenergetic functions (Matsuno-Yagi and Mukohata, Reference Matsuno-Yagi and Mukohata1977; Lindley and MacDonald, Reference Lindley and MacDonald1979; Lanyi and Weber, Reference Lanyi and Weber1980; Matsuno-Yagi and Mukohata, Reference Matsuno-Yagi and Mukohata1980; Mukohata and Kaji, Reference Mukohata and Kaji1981; Schobert and Lanyi, Reference Schobert and Lanyi1982).
In the early 1980s, a third rhodopsin-like protein was found in H. salinarum, which contributes to its phototaxis, and thus was named sensory rhodopsin (Bogomolni and Spudich, Reference Bogomolni and Spudich1982; Spudich and Spudich, Reference Spudich and Spudich1982; Spudich and Bogomolni, Reference Spudich and Bogomolni1984); this molecule did not pass ions, but instead triggered a nonionic signal transduction chain to control flagellar movement (Hoff et al., Reference Hoff, Jung and Spudich1997).
In the years since these early discoveries, a search throughout the tree of life for other such rhodopsin-like proteins in microbes has yielded a great many different versions, with different spectral sensitivities, kinetics, ion sensitivities, structures and internal mechanisms, and other properties, from diverse archaea and bacteria, and even eukaryotes such as fungi (Bieszke et al., Reference Bieszke, Braun, Bean, Kang, Natvig and Borkovich1999a, Reference Bieszke, Spudich, Scott, Borkovich and Spudich1999b). Some of these molecules, as noted above, such as a light-driven proton pump from H. sodomense (Chow et al., Reference Chow, Han, Dobry, Qian, Chuong, Li, Henninger, Belfort, Lin, Monahan and Boyden2010), and a light-driven chloride pump from N. pharaonis (Han and Boyden, Reference Han and Boyden2007; Zhang et al., Reference Zhang, Wang, Brauner, Liewald, Kay, Watzke, Wood, Bamberg, Nagel, Gottschalk and Deisseroth2007; Gradinaru et al., Reference Gradinaru, Zhang, Ramakrishnan, Mattis, Prakash, Diester, Goshen, Thompson and Deisseroth2010), have become widespread in neuroscience for light-driven neural silencing.
Curiously, even bacteriorhodopsin, the first microbial opsin to be discovered, could mediate sizable inhibitory photocurrents in cultured neurons, suggesting that perhaps the use of microbial rhodopsins to make neurons controllable by light could have begun years earlier, in principle (Chow et al., Reference Chow, Han, Dobry, Qian, Chuong, Li, Henninger, Belfort, Lin, Monahan and Boyden2010).
One of the most important discoveries that contributed to the development of optogenetics, was that specific rhodopsins mediated algal phototaxis, by converting light signals into fast ion channel currents (Foster et al., Reference Foster, Saranak, Patel, Zarilli, Okabe, Kline and Nakanishi1984; Harz and Hegemann, Reference Harz and Hegemann1991; Hegemann et al., Reference Hegemann, Gärtner and Uhl1991; Lawson et al., Reference Lawson, Zacks, Derguini, Nakanishi and Spudich1991; Takahashi et al., Reference Takahashi, Yoshihara, Watanabe, Kubota, Johnson, Derguini and Nakanishi1991; Sineshchekov et al., Reference Sineshchekov, Jung and Spudich2002). Algal phototaxis had been documented more than 150 years ago by Famintsyn, who described the effect of light intensity on the movement of the unicellular alga C. reinhardtii (Deisseroth and Hegemann, Reference Deisseroth and Hegemann2017; Salomé and Merchant, Reference Salomé and Merchant2019). One of the genes mediating this response in C. reinhardtii was found, upon expression in oocytes, to encode a light-gated proton channel, named channelrhodopsin-1 (Nagel et al., Reference Nagel, Ollig, Fuhrmann, Kateriya, Musti, Bamberg and Hegemann2002), and the other gene, upon expression in oocytes, HEK293, and BHK cells, was found to encode a nonspecific cation channel, named channelrhodopsin-2 (Nagel et al., Reference Nagel, Szellas, Huhn, Kateriya, Adeishvili, Berthold, Ollig, Hegemann and Bamberg2003). The latter molecule was able to mediate optical neural activation with single-spike precision (Boyden et al., Reference Boyden, Zhang, Bamberg, Nagel and Deisseroth2005) and is the most widespread molecule for neural activation with light. Since these papers, many new ion pumps and channels of many kinds, discussed in the next section, have been discovered, many with specialized and powerful applications in neuroscience.
In parallel to these discoveries, scientists and engineers were finding that these microbial rhodopsins could be genetically expressed in other organisms, both to achieve bioengineering goals, as well as to facilitate their study. One early study expressed bacteriorhodopsin in E. coli, to facilitate its study, although expression was poor (Dunn et al., Reference Dunn, Hackett, McCoy, Chao, Kimura and Khorana1987), and codon optimization and signal sequence addition had to be performed to improve yield (Karnik et al., Reference Karnik, Nassal, Doi, Jay, Sgaramella and Khorana1987).
Later studies showed that bacteriorhodopsin could be expressed in eukaryotic cells. One such study expressed bacteriorhodopsin in yeast (Hildebrandt et al., Reference Hildebrandt, Ramezani-Rad, Swida, Wrede, Grzesiek, Primke and Büldt1989), and found that the protein was able to pump protons across the plasma membrane, out of the cell (Hildebrandt et al., Reference Hildebrandt, Fendler, Heberle, Hoffmann, Bamberg and Büldt1993). Targeted expression of bacteriorhodopsin to the mitochondria of yeast enabled them to rely less on sugar for metabolism, equipping the yeast with a rudimentary form of photosynthesis (Hoffmann et al., Reference Hoffmann, Hildebrandt, Heberle and Büldt1994) – perhaps one of the first applications of microbial rhodopsins to a bioengineering goal.
Regarding vertebrate cells: frog oocytes expressed the gene for bacteriorhodopsin, and exhibited light-driven currents (Nagel et al., Reference Nagel, Möckel, Büldt and Bamberg1995); this facilitated the use of voltage clamp and patch clamp methods to characterize the photocurrents. Bacteriorhodopsin could be also expressed in cultured mammalian cells, using the human HEK293 cell line, where it exhibited excellent photocurrents (Geibel et al., Reference Geibel, Friedrich, Ormos, Wood, Nagel and Bamberg2001); this study also showed that membrane expression could be boosted in these animal cells by appending a targeting sequence. These studies led to many downstream experiments in a variety of cell types, both revealing fundamental biophysical properties of rhodopsins, as well as paving the way for broader and broader application of rhodopsins toward different engineering goals.
Structure and biophysics of microbial rhodopsins
Opsin classification and structure
Microbial rhodopsins, both natural and engineered, exhibit a variety of structural and biophysical properties that help them mediate powerful, specific neural electrical activity control in response to light (Figure 2). In the remainder of this review, we go over the opsin classes and their structural properties, followed by sections on their photocycles, their photocurrent magnitudes and kinetics, their action spectra, and their ion selectivities, both diving into the biophysical mechanisms underlying these properties, and how these properties fit well with urgent neuroscience needs.
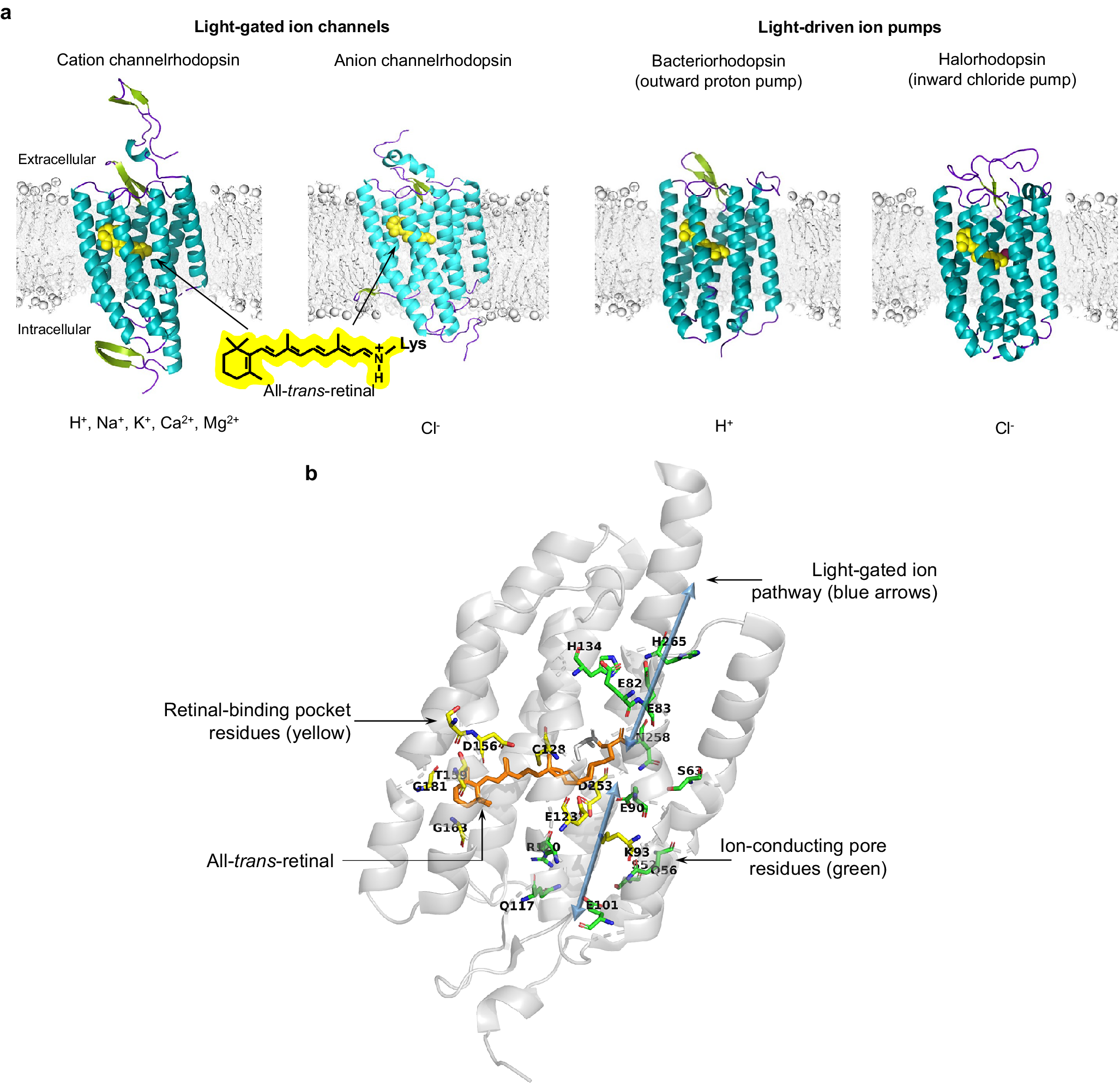
Figure 2. 3D protein structure and chromophore-protein interactions of rhodopsins. (a, from left to right) 3D protein structures of single subunits and respective conducted ions for the C1C2 cation channelrhodopsin (PDB 3UG9), the GtACR1 anion channelrhodopsin (PDB 6CSM), the archaerhodopsin-2 outward proton pump (PDB 2EI4), and the N. pharaonis inward chloride pump (PDB 3A7K). (b) Key residues in the ChR2 channelrhodopsin (PDB 6EID).
Microbial rhodopsins, also called type I rhodopsins (as opposed to the type II rhodopsins found in animals, which are G-protein coupled), are found in bacteria, archaea, algae, and other species, where they mediate light-driven energy conversion or light-driven sensory transduction processes (Govorunova et al., Reference Govorunova, Sineshchekov, Li and Spudich2017). Based on their biophysical properties, the microbial rhodopsins used in neuroscience for mediating the control of neural electrical activity with light can be divided into four major groups: light-driven outward proton pumps (also referred to as bacteriorhodopsins or BRs), light-driven inward chloride pumps (also referred to as halorhodopsins or HRs), light-activated cation channels (often referred to as channelrhodopsins, ChRs, or more recently cation channelrhodopsins, CCRs), and light-activated anion channels (often referred to as anion channelrhodopsins or ACRs).
In addition, a fifth group of microbial rhodopsins, represented by recently discovered potassium-selective channelrhodopsins (referred to as kalium channelrhodopsins, KCRs) (Govorunova et al., Reference Govorunova, Gou, Sineshchekov, Li, Lu, Wang, Brown, St-Pierre, Xue and Spudich2022; Vierock et al. Reference Vierock, Peter, Grimm, Andrey Rozenberg, Tillert, Scalise, Casini, Augustin, Tanese, Forget, Peyronnet, Schneider-Warme, Emiliani, Béjà and Hegemann2022), has emerged. Such rhodopsins pass cations, but in contrast to other light-gated cation channels, are outward-passing channels, and thus cause neural silencing effects. KCRs hold great promise for neuroscience applications, and as they are explored, validated, and optimized in different contexts, they may find many powerful uses in neuroscience (Govorunova et al., Reference Govorunova, Sineshchekov and Spudich2023). For the purposes of this review, which focuses on biophysics of neural control, we focus on the four major groups of microbial rhodopsins that have been most thoroughly biophysically characterized.
Despite distinct mechanisms of ion transport and varying biophysical characteristics, all microbial rhodopsins share a relatively high overall amino acid similarity (Spudich et al., Reference Spudich, Yang, Jung and Spudich2000; Man et al., Reference Man, Wang, Gazalah Sabehi, Post, Massana, Spudich, Spudich and Béjà2003; Song and Gunner, Reference Song and Gunner2014), ranging from 25–80% homology, as well as a highly conserved overall 3D structure comprising seven α-helix transmembrane domains (Pebay-Peyroula et al., Reference Pebay-Peyroula, Rummel, Rosenbusch and Landau1997; Kolbe et al., Reference Kolbe, Besir, Essen and Oesterhelt2000; Kato et al., Reference Kato, Zhang, Yizhar, Ramakrishnan, Nishizawa, Hirata, Ito, Aita, Tsukazaki, Hayashi, Hegemann, Maturana, Ishitani, Deisseroth and Nureki2012). The core of an opsin comprises ~250–320 amino acids, and incorporates the obligate co-factor all-trans-retinal, which serves as the photosensitive moiety (Figure 2). Retinal attaches to a specific lysine side chain on the opsin protein, autocatalytically via a protonated Schiff base linkage, forming the functional opsin protein, termed rhodopsin. The N-terminal domain of rhodopsins is exposed to the extracellular space and the C-terminal domain is located intracellularly, and is often fused to a fluorescent protein for opsin expression visualization.
Comparative analysis of channelrhodopsins and ion pumps revealed several distinct structural features of these two classes of optogenetic tool. First, wild-type ChRs, but not wild-type light-driven pumps, harbor an intracellular signaling domain (Nagel et al., Reference Nagel, Szellas, Huhn, Kateriya, Adeishvili, Berthold, Ollig, Hegemann and Bamberg2003) which contributes to subcellular localization and signaling function in native organisms (Mittelmeier et al., Reference Mittelmeier, Boyd, Lamb and Dieckmann2011). This intracellular domain is not required for photocurrent generation, and is usually removed during biophysical investigations of photocurrent (and perhaps replaced with a fluorescent protein to facilitate visualization during heterologous expression) (Nagel et al., Reference Nagel, Ollig, Fuhrmann, Kateriya, Musti, Bamberg and Hegemann2002, Reference Nagel, Szellas, Huhn, Kateriya, Adeishvili, Berthold, Ollig, Hegemann and Bamberg2003). Determination of the first crystal structures for the wild-type and chimeric channelrhodopsins, ChR2 and C1C2, respectively, revealed a dimeric oligomerization state (Müller et al., Reference Müller, Bamann, Bamberg and Kühlbrandt2011; Kato et al., Reference Kato, Zhang, Yizhar, Ramakrishnan, Nishizawa, Hirata, Ito, Aita, Tsukazaki, Hayashi, Hegemann, Maturana, Ishitani, Deisseroth and Nureki2012), which has been seen for other cation and anion ChRs with solved crystal structure, such as C1C2 (with improved resolution) (Volkov et al., Reference Volkov, Kovalev, Polovinkin, Borshchevskiy, Bamann, Astashkin, Marin, Popov, Balandin, Willbold, Büldt, Bamberg and Gordeliy2017), C1Chrimson (Oda et al., Reference Oda, Vierock, Oishi, Rodriguez-Rozada, Taniguchi, Keitaro Yamashita, Nishizawa, Hegemann and Nureki2018), GtACR1 (Kim et al., Reference Kim, Kato, Yamashita, Ito, Inoue, Ramakrishnan, Fenno, Evans, Paggi, Dror, Kandori, Kobilka and Deisseroth2018), and iC++ (Kato et al., Reference Kato, Kim, Paggi, Evans, Allen, Richardson, Inoue, Ito, Ramakrishnan, Fenno, Yamashita, Hilger, Lee, Berndt, Shen, Kandori, Dror, Kobilka and Deisseroth2018); the newly developed red-shifted ChR ChRmine was reported to form trimers (Kishi et al., Reference Kishi, Kim, Fukuda, Inoue, Kusakizako, Wang, Ramakrishnan, EFX, Thadhani, Paggi, Matsui, Yamashita, Nagata, Konno, Quirin, Lo, Benster, Uemura, Liu, Shibata, Nomura, Iwata, Nureki, Dror, Inoue, Deisseroth and Kato2022), perhaps more similar to light-driven ion pumps (Kishi et al., Reference Kishi, Kim, Fukuda, Inoue, Kusakizako, Wang, Ramakrishnan, EFX, Thadhani, Paggi, Matsui, Yamashita, Nagata, Konno, Quirin, Lo, Benster, Uemura, Liu, Shibata, Nomura, Iwata, Nureki, Dror, Inoue, Deisseroth and Kato2022). Indeed, BRs and HRs mostly exist in trimers in native membrane environments (Essen et al., Reference Essen, Siegert, Lehmann and Oesterhelt1998; Sasaki et al., Reference Sasaki, Kubo, Kikukawa, Kamiya, Aizawa, Kawano, Kamo and Demura2009; Shibata et al., Reference Shibata, Yamashita, Uchihashi, Kandori and Ando2010, Reference Shibata, Inoue, Ikeda, Konno, Singh, Kataoka, Abe-Yoshizumi, Kandori and Uchihashi2018), although it was shown that the functional unit responsible for the ion transport photocycle is the monomeric form (Dencher and Heyn, Reference Dencher and Heyn1979; Grzesiek and Dencher, Reference Grzesiek and Dencher1988). Oligomerization of BRs improves their structural stability and increases incorporation of all-trans-retinal (Dencher et al., Reference Dencher, Kohl and Heyn1983; Brouillette et al., Reference Brouillette, McMichens, Stern and Khorana1989); trimer–trimer interactions may also facilitate the full natural photo-reaction pathway (Yamashita et al., Reference Yamashita, Inoue, Shibata, Uchihashi, Sasaki, Kandori and Ando2013). Due to the oligomeric state of rhodopsins, C-terminal fusions to monomeric fluorescent proteins may help minimize disruption of opsin localization and function, in neuroscience contexts.
The high-resolution crystal structures of microbial rhodopsins have provided much insight into ion conduction and transport specificity due to specific amino acid configurations, as well as chromophore-amino acid interactions that regulate the colors of light that best drive opsin function (Figure 2b). For example, crystal structures of bacteriorhodopsins and halorhodopsins revealed molecular details of ion transport pathways and mechanisms, including structures of intermediate states after light absorption, and key amino acids (and key bound water molecules) that bind to, transport, and release ions in a directional fashion along the pathway through the protein that crosses the membrane (Luecke et al., Reference Luecke, Richter and Lanyi1998, Reference Luecke, Schobert, Richter, Cartailler and Lanyi1999a, Reference Luecke, Schobert, Richter, Cartailler and Lanyi1999b; Facciotti et al., Reference Facciotti, Rouhani, Burkard, Betancourt, Downing, Rose, McDermott and Glaeser2001; Lanyi and Luecke, Reference Lanyi and Luecke2001; Patzelt et al., Reference Patzelt, Simon, TerLaak, Kessler, Kühne, Schmieder, Oesterhelt and Oschkinat2002; Enami et al., Reference Enami, Yoshimura, Murakami, Okumura, Ihara and Kouyama2006; Kouyama et al., Reference Kouyama, Kanada, Takeguchi, Narusawa, Murakami and Ihara2010; Song and Gunner, Reference Song and Gunner2014; Kouyama et al., Reference Kouyama, Kawaguchi, Nakanishi, Kubo and Murakami2015; Mous et al., Reference Mous, Gotthard, Ehrenberg, Sen, Weinert, Philip, James, Nass, Furrer, Kekilli, Ma, Brünle, Casadei, Martiel, Dworkowski, Gashi, Skopintsev, Wranik, Knopp, Panepucci, Panneels, Cirelli, Ozerov, GFX, Wang, Milne, Standfuss, Schapiro, Heberle and Nogly2022). 3D structures of cation and anion ChRs such as C1C2 (with improved resolution over the first reported structure) (Volkov et al., Reference Volkov, Kovalev, Polovinkin, Borshchevskiy, Bamann, Astashkin, Marin, Popov, Balandin, Willbold, Büldt, Bamberg and Gordeliy2017), the cation channelrhodopsin C1Chrimson (chimera of ChR1 and CsChrimson) (Oda et al., Reference Oda, Vierock, Oishi, Rodriguez-Rozada, Taniguchi, Keitaro Yamashita, Nishizawa, Hegemann and Nureki2018), the natural ACR called GtACR1 (Kim et al., Reference Kim, Kato, Yamashita, Ito, Inoue, Ramakrishnan, Fenno, Evans, Paggi, Dror, Kandori, Kobilka and Deisseroth2018; Li et al., Reference Li, Huang, Govorunova, Schafer, Sineshchekov, Wang, Zheng and Spudich2019, Reference Li, Huang, Govorunova, Sineshchekov, Wang, Zheng and Spudich2021), the engineered ACR iC++ (Kato et al., Reference Kato, Kim, Paggi, Evans, Allen, Richardson, Inoue, Ito, Ramakrishnan, Fenno, Yamashita, Hilger, Lee, Berndt, Shen, Kandori, Dror, Kobilka and Deisseroth2018), and the red-shifted engineered cation channelrhodopsin ChRmine (Kishi et al., Reference Kishi, Kim, Fukuda, Inoue, Kusakizako, Wang, Ramakrishnan, EFX, Thadhani, Paggi, Matsui, Yamashita, Nagata, Konno, Quirin, Lo, Benster, Uemura, Liu, Shibata, Nomura, Iwata, Nureki, Dror, Inoue, Deisseroth and Kato2022) revealed molecular determinants of rhodopsins’ kinetics, spectral tuning, and ion selectivity. High-resolution structures of rhodopsins are guiding the rational design of novel optogenetic tools with altered biophysical properties, enabling tools customized for specialized needs in neuroscience (Kaneko et al., Reference Kaneko, Inoue, Kojima, Kandori and Sudo2017). In addition, genomic search and molecular mutant screening are enabling the identification of novel rhodopsins and the tuning of properties of rhodopsins, including photocurrent (Figure 3), spectral tuning (Figure 4), light sensitivity (Figure 5), kinetics, and many other features (Figure 6).
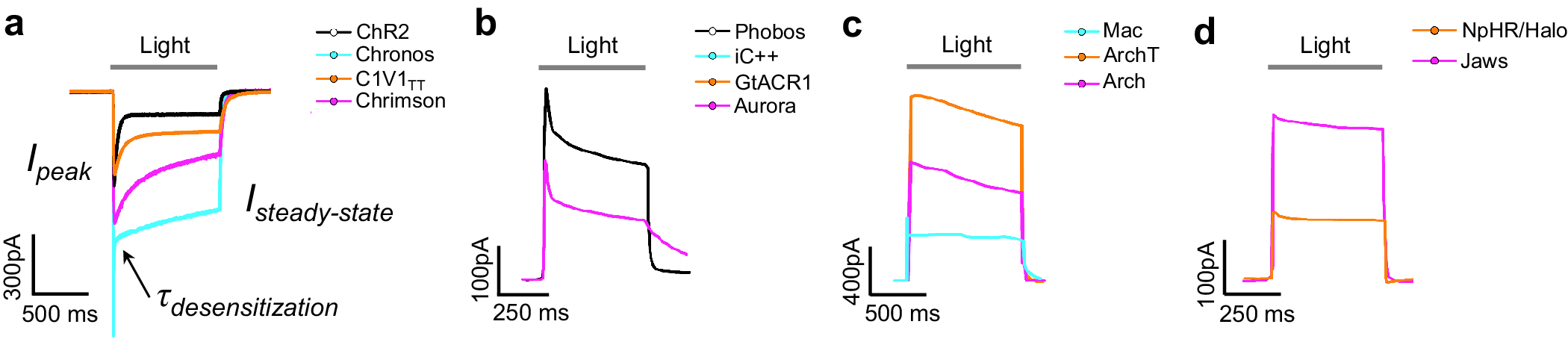
Figure 3. Photocurrent traces of representative rhodopsins. (a) Photocurrent traces of the ChR2, Chronos, C1V1TT, and Chrimson cation channelrhodopsins showing peak photocurrent (I peak), steady-state photocurrent (I steady-state), and desensitization kinetics (τ desensitization). (b) Photocurrent traces of the Phobos, iC++, GtACR1, and Aurora anion channelrhodopsins (measured in HEK cells). (c) Photocurrent traces of the Mac, ArchT, and Arch outward proton pumps (measured in cultured neurons). (d) Photocurrent traces of the NpHR/Halo and Jaws inward chloride pumps (measured in cultured neurons). Traces are recorded in cultured cells under saturating light powers near respective peak wavelengths of corresponding rhodopsins at holding potential of −70 mV. Data from Chuong et al. (Reference Chuong, Miri, Busskamp, Matthews, Acker, Sørensen, Young, Klapoetke, Henninger, Kodandaramaiah, Ogawa, Ramanlal, Bandler, Allen, Forest, Chow, Han, Lin, Tye, Roska, Cardin and Boyden2014), Klapoetke et al. (Reference Klapoetke, Murata, Kim, Pulver, Birdsey-Benson, Cho, Morimoto, Chuong, Carpenter, Tian, Wang, Xie, Yan, Zhang, Chow, Surek, Melkonian, Jayaraman, Constantine-Paton, Wong and Boyden2014), Govorunova et al. (Reference Govorunova, Sineshchekov, Janz, Liu and Spudich2015), and Wietek et al. (Reference Wietek, Rodriguez-Rozada, Tutas, Tenedini, Grimm, Oertner, Soba, Hegemann and Wiegert2017).
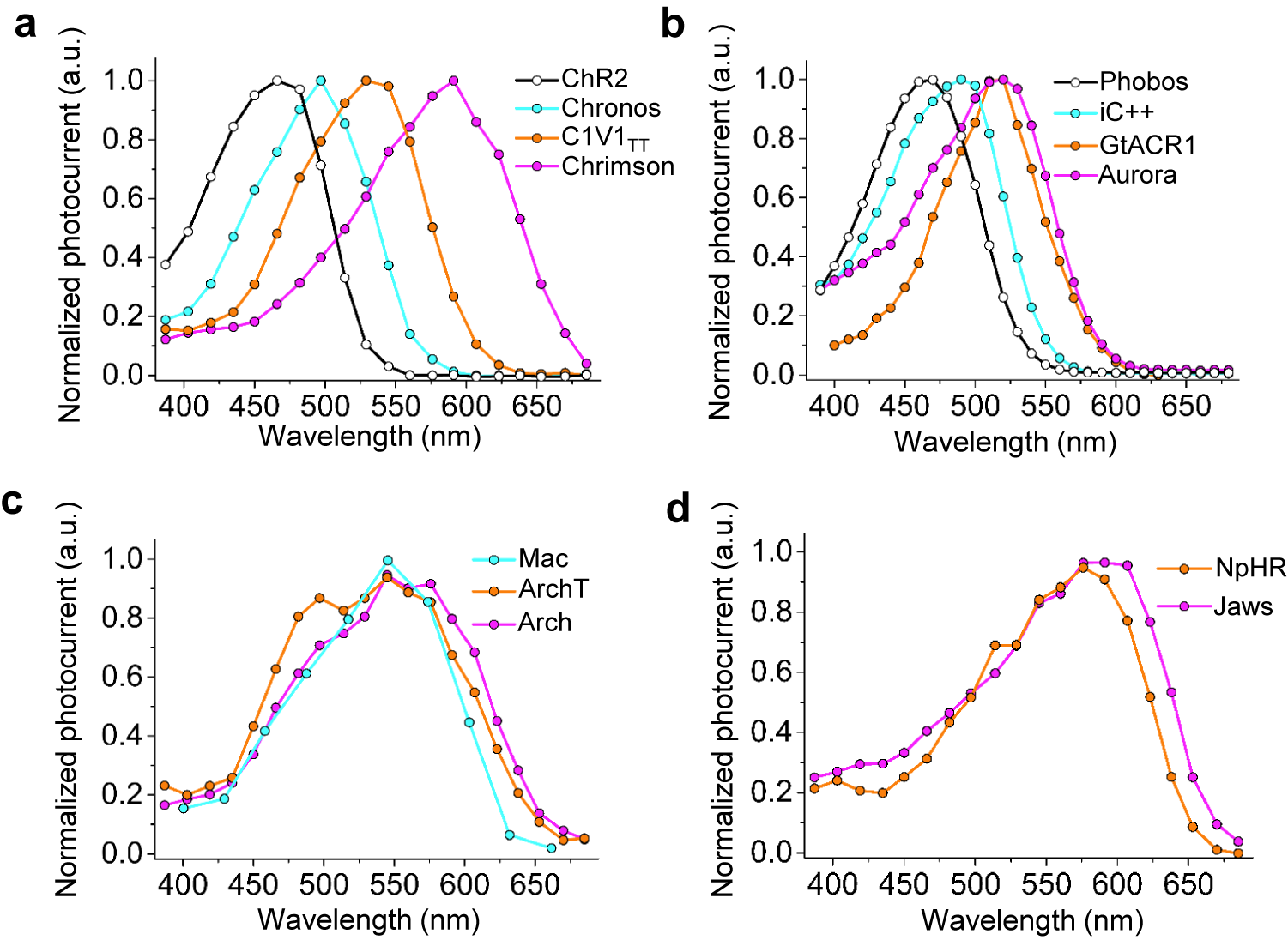
Figure 4. Action spectra of representative rhodopsins. (a) Action spectra of the ChR2, Chronos, C1V1TT, and Chrimson cation channelrhodopsins (measured in HEK cells). (b) Action spectra of the Phobos, iC++, GtACR1, and Aurora anion channelrhodopsins (measured in HEK cells). (c) Action spectra of the Mac, ArchT, and Arch outward proton pumps (measured in cultured neurons). (d) Action spectra of the NpHR and Jaws inward chloride pumps (measured in cultured neurons). Data from Chuong et al. (Reference Chuong, Miri, Busskamp, Matthews, Acker, Sørensen, Young, Klapoetke, Henninger, Kodandaramaiah, Ogawa, Ramanlal, Bandler, Allen, Forest, Chow, Han, Lin, Tye, Roska, Cardin and Boyden2014), Klapoetke et al. (Reference Klapoetke, Murata, Kim, Pulver, Birdsey-Benson, Cho, Morimoto, Chuong, Carpenter, Tian, Wang, Xie, Yan, Zhang, Chow, Surek, Melkonian, Jayaraman, Constantine-Paton, Wong and Boyden2014), Govorunova et al. (Reference Govorunova, Sineshchekov, Janz, Liu and Spudich2015), and Wietek et al. (Reference Wietek, Rodriguez-Rozada, Tutas, Tenedini, Grimm, Oertner, Soba, Hegemann and Wiegert2017).
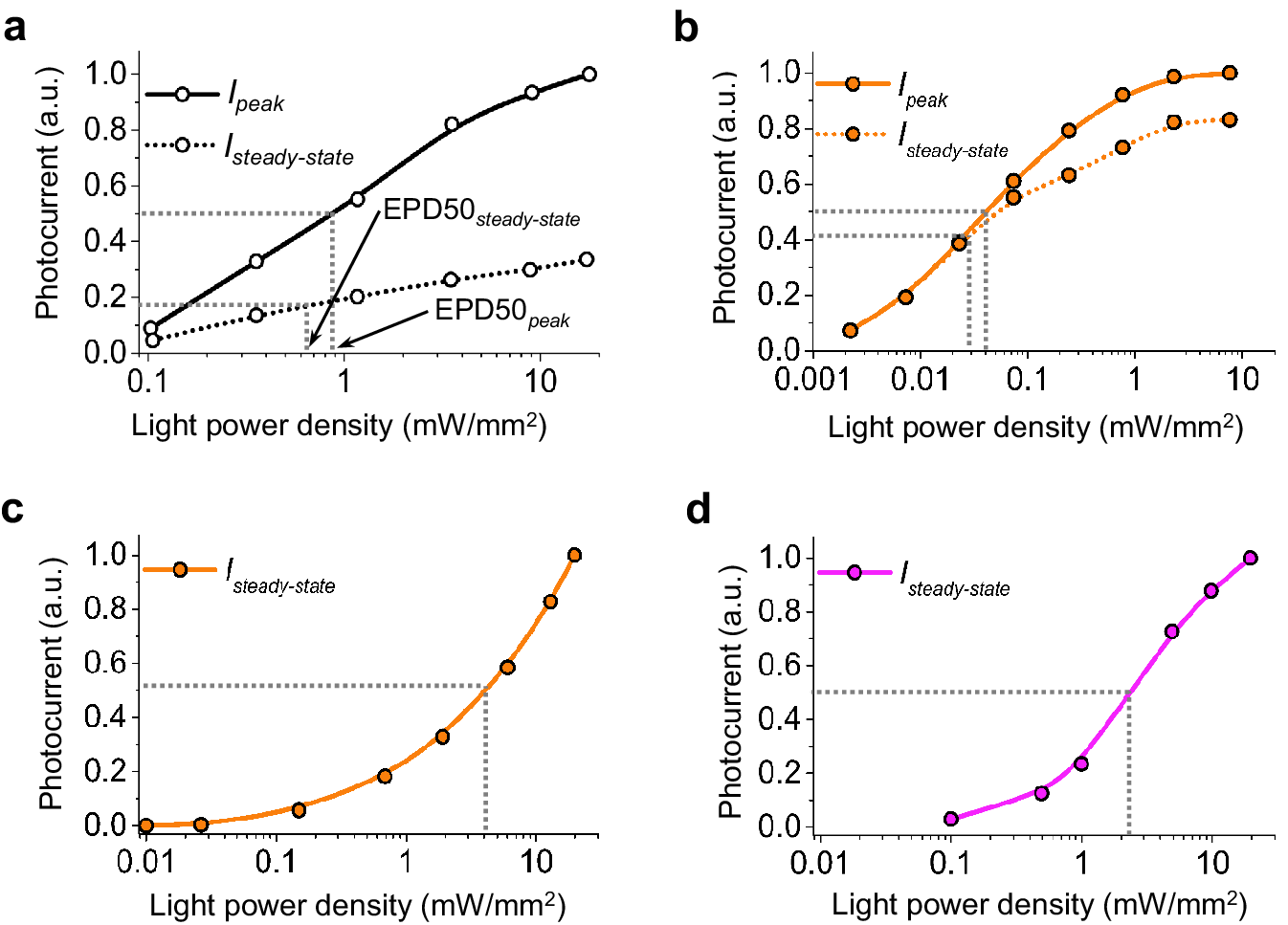
Figure 5. Light sensitivity of representative rhodopsins. (a–d) Peak (solid line) and steady-state (dashed line) photocurrents across light intensities for (a) ChR2 (measured in cultured neurons), (b) GtACR1 (measured in HEK cells), (c) ArchT (measured in cultured neurons), and (d) Jaws (measured in cultured neurons). Data from Chuong et al. (Reference Chuong, Miri, Busskamp, Matthews, Acker, Sørensen, Young, Klapoetke, Henninger, Kodandaramaiah, Ogawa, Ramanlal, Bandler, Allen, Forest, Chow, Han, Lin, Tye, Roska, Cardin and Boyden2014), Klapoetke et al. (Reference Klapoetke, Murata, Kim, Pulver, Birdsey-Benson, Cho, Morimoto, Chuong, Carpenter, Tian, Wang, Xie, Yan, Zhang, Chow, Surek, Melkonian, Jayaraman, Constantine-Paton, Wong and Boyden2014), Govorunova et al. (Reference Govorunova, Sineshchekov, Janz, Liu and Spudich2015), and Wietek et al. (Reference Wietek, Rodriguez-Rozada, Tutas, Tenedini, Grimm, Oertner, Soba, Hegemann and Wiegert2017).
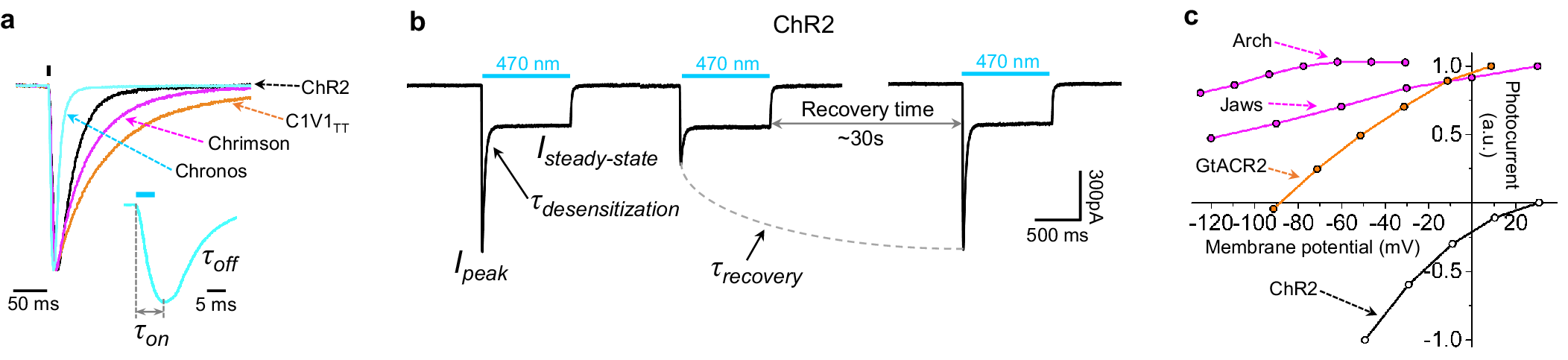
Figure 6. Biochemical and biophysical properties of representative rhodopsins. (a) Photocurrent traces generated by 5-ms illumination near peak wavelength of indicated rhodopsins expressed in cultured neurons. Traces are normalized to facilitate the comparison of photocurrent kinetics. (b) Traces of photocurrent recovery kinetics for ChR2 measured in cultured neurons. (c) Photocurrent-voltage relationships curves for ChR2, GtACR2, Arch, and Jaws, measured in HEK cells. Data from Chuong et al. (Reference Chuong, Miri, Busskamp, Matthews, Acker, Sørensen, Young, Klapoetke, Henninger, Kodandaramaiah, Ogawa, Ramanlal, Bandler, Allen, Forest, Chow, Han, Lin, Tye, Roska, Cardin and Boyden2014), Klapoetke et al. (Reference Klapoetke, Murata, Kim, Pulver, Birdsey-Benson, Cho, Morimoto, Chuong, Carpenter, Tian, Wang, Xie, Yan, Zhang, Chow, Surek, Melkonian, Jayaraman, Constantine-Paton, Wong and Boyden2014), Govorunova et al. (Reference Govorunova, Sineshchekov, Janz, Liu and Spudich2015), and Wietek et al. (Reference Wietek, Rodriguez-Rozada, Tutas, Tenedini, Grimm, Oertner, Soba, Hegemann and Wiegert2017).
Photocycle
Upon photon absorption, the retinal chromophore of an opsin undergoes isomerization, initiating a series of functional and conformational changes in the protein, also known as the photocycle (Stehfest and Hegemann, Reference Stehfest and Hegemann2010; Schneider et al., Reference Schneider, Grimm and Hegemann2015). These light-induced protein conformational changes result in ion transport across the membrane in which the protein is embedded, measured electrophysiologically as photocurrent, either by opening an ion-permeable pore in rhodopsin, thus allowing multiple ions to passively cross the membrane (bidirectionally, governed by the voltage across the membrane, the concentration of ions on either side of the membrane, and any rectification or other intrinsic properties of the rhodopsin) per absorbed photon, or by actively pumping ions, uni-directionally translocating one ion per absorbed photon in a fashion that is less dependent on ion concentration and membrane voltage. The opsin photocycle involves multiple, usually short-lived (e.g., lasting microseconds to milliseconds) intermediate states characterized by different conformations of the retinal chromophore, different protein conformations, and different interactions between the retinal and local amino acids. The intermediates are traditionally named for their absorption peaks, as determined by standard absorption spectroscopy. A halorhodopsin, for example, will typically start out in a state where its absorption peak is in the yellow range (i.e., ~580 nm), and upon illumination it will rapidly change into a conformation that has an absorption peak of 600 nm, followed by conformations with absorption peaks of 520, 640, and 565 nm, followed by a reversion back to a conformation with peak absorption of 580 nm (Essen, Reference Essen2002).
Microbial rhodopsins have closed photocycles, that is, they end up in the same state as they started, which is one reason they are useful tools in biology. In contrast, the type II rhodopsins of mammalian photoreceptors end in a state that is covalently different from their initial state, requiring significant cellular machinery for their recycling into an active form. It should be noted though that not all conformational changes are associated with a spectroscopic shift, so such descriptions are an approximation, albeit a useful one. The initial transition is extremely fast, taking less than a nanosecond to occur. During the latter conformational changes, key sets of amino acids, which serve as ion binding sites, capable of strong electrostatic interactions with target ions to assist with their transport, will change conformation, causing the ion of interest to be handed off from site to site throughout the protein, with sites changing ion affinity as appropriate, so that the ion eventually traverses the membrane (from the extracellular side to the cytoplasmic side, for a halorhodopsin; in the other direction, for a bacteriorhodopsin). The initial uptake of an ion, and the final release of the ion into the environment, are governed by passive diffusion from/to the environment, so availability of appropriate ions in sufficient concentrations is essential.
Photocurrent magnitude and kinetics
Whereas light-driven ion pumps transport one ion per photon absorbed, light-driven ion channels can transport multiple ions per photon absorbed. The effectiveness of light in inducing voltage changes in a target neuron is in significant part determined by the number of ions that can be translocated by a rhodopsin across the membrane per unit of time, which is defined as the unitary photocurrent, times the number of functional proteins in the membrane, which is a measure of protein-membrane expression (including successful membrane trafficking, protein folding, and retinal incorporation, amongst other key factors). For a light-gated ion channel, it is challenging to obtain the unitary photocurrent of a single rhodopsin molecule, because its ion conductance is several orders of magnitude lower than that of the voltage and ligand-gated ion channels typically studied in neuroscience (Baumgarten et al., Reference Baumgarten, Dudley, Rogart and Fozzard1995; Picones et al., Reference Picones, Keung and Timpe2001; Doering et al., Reference Doering, Hamid, Simms, McRory and Zamponi2005), and below the limit for direct measurements with state-of-the-art methods like single channel patch clamp. Thus, various research groups have used different methods to estimate the ion conductance of a single light-gated ion channel, its unitary conductance (Nagel et al., Reference Nagel, Szellas, Huhn, Kateriya, Adeishvili, Berthold, Ollig, Hegemann and Bamberg2003; Feldbauer et al., Reference Feldbauer, Zimmermann, Pintschovius, Spitz, Bamann and Bamberg2009; Lin et al., Reference Lin, Lin, Steinbach and Tsien2009; Kleinlogel et al., Reference Kleinlogel, Feldbauer, Dempski, Fotis, Wood, Bamann and Bamberg2011; Govorunova et al., Reference Govorunova, Sineshchekov, Li, Janz and Spudich2013). Due to the different methods employed by the various groups, the estimated unitary conductance reported for the same rhodopsin variant can vary, sometimes by an order of magnitude, between reports (Harz et al., Reference Harz, Nonnengasser and Hegemann1992; Nagel et al., Reference Nagel, Szellas, Huhn, Kateriya, Adeishvili, Berthold, Ollig, Hegemann and Bamberg2003; Lin et al., Reference Lin, Lin, Steinbach and Tsien2009). However, over time, general consensuses can emerge; for example, the unitary conductance of ChR2, one of the most widely used cation channelrhodopsins, estimated by stationary noise analysis, is in the range of 30–40 fS (Feldbauer et al., Reference Feldbauer, Zimmermann, Pintschovius, Spitz, Bamann and Bamberg2009; Govorunova et al., Reference Govorunova, Sineshchekov, Li, Janz and Spudich2013), which corresponds to translocation of 10–14 ions per molecule during one typical photocycle, equating to approximately 103–104 ions per molecule per second. Overall, the unitary conductance of rhodopsins can vary, across molecules, over an order of magnitude or more: for example, the PsChR channelrhodopsin has threefold higher ion conductance versus that of ChR2 (Govorunova et al., Reference Govorunova, Sineshchekov, Li, Janz and Spudich2013), and the largest unitary conductance among rhodopsins was perhaps demonstrated by the natural anion channelrhodopsin GtACR2, reaching ~600 fS (Govorunova et al., Reference Govorunova, Sineshchekov, Janz, Liu and Spudich2015).
Despite their low unitary conductances, ChRs can efficiently drive actional potentials in neuroscience experiments, since they are only required to depolarize neural membranes above the action potential threshold, which can be quite low. For example, activation of only ~170,000 ChR2 molecules (or ~ 240 molecules/μm2 in a soma of 15 μm diameter) could be sufficient to evoke an action potential, whereas a typical opsin expression level might be 100–500 molecules/μm2. In more detail: assuming linearity, and no changes in membrane resistance during depolarization: (1) assume a ballpark neural membrane resistance of 50 MΩ; (2) a ~15 mV depolarization is sufficient to cross action potential threshold; (3) the voltage or ion driving force experienced by the ChR2 molecule is 0–(−60 mV) = 60 mV; (4) the unitary conductance of ChR2 is 30fS; (5) then, the current needed to depolarize the neuron by 15 mV would be I = 15 mV/50 MΩ = 300 pA = 3 × 10−10A; (6) each molecule of ChR2 is capable of carrying g × V = 30 fS × 60 mV = 1.8 fA = 1.8 × 10−15 A; (7) N = 300 pA/1.8 fA = ~170,000 molecules or 240 molecules/μm2; (8) since the driving force experienced by ChR2 will be reduced by ~25% upon 15 mV depolarization, we can calculate an estimated bound on the maximum number of ChR2 molecules needed, by increasing the final channel count by 25%, to 75 molecules/μm2. Given an estimated surface area of the neuron soma of ~700 μm2 (soma of 15 μm), the average expression level of rhodopsins in neurons could be estimated as ~100–500 molecules/μm2, sufficient for neural control, based on the calculations above. However, it should be taken into account that the maximum photocurrent is achieved at saturating light power, which depends on the light sensitivity of the opsin (considered in more detail below); therefore the actual probability of eliciting a spike will also be determined by the illumination intensity. Similar calculations can be also applied to light-driven pumps, which have higher driving force (indeed, being pumps, they can transport ions even against a gradient), and therefore will provide more constant photocurrent amidst voltage fluctuations; however, the light sensitivity of a pump is typically severalfold lower than that of a channel, and thus typically will require higher illumination intensity.
Due to the difficulty in measuring it, the unitary conductance is often not reported for an opsin when it is being characterized for its performance as a neuroscience tool, but rather the total photocurrent generated by all functional opsin molecules in a single cell, is measured and reported (Lin et al., Reference Lin, Lin, Steinbach and Tsien2009; Mattis et al., Reference Mattis, Tye, Ferenczi, Ramakrishnan, O’Shea, Prakash, Gunaydin, Hyun, Fenno, Gradinaru, Yizhar and Deisseroth2011; Klapoetke et al., Reference Klapoetke, Murata, Kim, Pulver, Birdsey-Benson, Cho, Morimoto, Chuong, Carpenter, Tian, Wang, Xie, Yan, Zhang, Chow, Surek, Melkonian, Jayaraman, Constantine-Paton, Wong and Boyden2014). Depending on the direction of the photocurrent generated under physiological conditions, rhodopsins can be classified into two major groups. The first group is represented by cation channelrhodopsins (CCRs) that generate inward-directed photocurrents carried by protons and cations, inducing depolarization of membrane potentials at the cell body, at neuron potentials from −80 to −60 mV. One exception to this is the recent discovery of channelrhodopsins with high potassium conductance, which conduct K+ outwards upon light gating, and thus have the opposite physiological effect, hyperpolarization rather than depolarization. In this review, we will thus call the older, H+/Na + -conducting CCRs depolarizing CCRs, and the new K + -conducting CCR a hyperpolarizing CCR or KCRs (Govorunova et al., Reference Govorunova, Sineshchekov and Spudich2023). In the second group, anion ChRs and light-driven ion pumps generate outward-directed photocurrents, thus causing inhibition of neural depolarization, or hyperpolarization.
ChRs and light-driven ion pumps exhibit distinct photocurrent profiles, due to their different mechanisms of ion translocation (Figure 3). The illumination of a ChR with a light pulse typically evokes a rapid rise in photocurrent, until it reaches a peak current (I peak), which then decays (in a fashion that can be often modeled by a biexponential decay) to a steady-state photocurrent (I steady-state) in a process denoted inactivation or desensitization. The kinetics of ChR photocurrent can be modeled by a four-state electrophysiological photocycle model (Nikolic et al., Reference Nikolic, Grossman, Grubb, Burrone, Toumazou and Degenaar2009). After rhodopsins are in the desensitized state, evoked photocurrents will be below the peak photocurrent, unless the ChR is allowed to recover in the dark, which can take several seconds or longer, depending on the opsin under consideration (Figure 6). Due to the need for prolonged illumination in many optogenetic applications, for example in the study of neural dynamics and behavior, an important parameter defining the performance of a rhodopsin from a neuroscience perspective is the ability to generate stable photocurrent responses over behaviorally relevant timescales. Photocurrent stability depends on steady-state/peak photocurrent ratio (I steady-state/I peak), desensitization kinetics (τ desensitization), and recovery kinetics from desensitization in darkness (τ recovery; Figures 3 and 6). The steady-state/peak photocurrent ratio represents the amount of photocurrent that persists during extended illumination, while τ desensitization and τ recovery correspond, respectively, to the rate of reduction and the rate of recovery of photocurrent to I peak when in the dark. Based upon these parameters, a rhodopsin possessing higher I steady-state/I peak, slower τ desensitization, and faster τ recovery, will generate more stable photocurrent during extended illumination periods, and would thus likely be more preferable – all else being equal – for a typical optogenetic experiment.
For example, based on photocurrent measurements performed in mammalian cells, ChR2 exhibits a > 70% drop in photocurrent within ~60 ms of illumination and requires about 30 seconds in darkness to completely restore its photocurrent (Lin et al., Reference Lin, Lin, Steinbach and Tsien2009; Lin, Reference Lin2011; Mattis et al., Reference Mattis, Tye, Ferenczi, Ramakrishnan, O’Shea, Prakash, Gunaydin, Hyun, Fenno, Gradinaru, Yizhar and Deisseroth2011; Figure 6). As a result, the probability of driving action potentials during a long train of light pulses quickly decreases for ChR2 at modest light power (2 mW/mm2), because the peak photocurrent rapidly declines and not much steady-state current is being elicited at this low power (Lin et al., Reference Lin, Lin, Steinbach and Tsien2009; Mattis et al., Reference Mattis, Tye, Ferenczi, Ramakrishnan, O’Shea, Prakash, Gunaydin, Hyun, Fenno, Gradinaru, Yizhar and Deisseroth2011; Figure 5a). However, it should be noted that the reliability of eliciting spikes during sustained light pulse trains, can be significantly improved at higher light intensities (20 mW/mm2) due to the higher contribution of steady-state photocurrent at these higher powers (Mattis et al., Reference Mattis, Tye, Ferenczi, Ramakrishnan, O’Shea, Prakash, Gunaydin, Hyun, Fenno, Gradinaru, Yizhar and Deisseroth2011; Figure 5a). The ChR2 mutant CatCh (Kleinlogel et al., Reference Kleinlogel, Feldbauer, Dempski, Fotis, Wood, Bamann and Bamberg2011) and the chimera rhodopsins C1V1TT (Yizhar et al., Reference Yizhar, Fenno, Prigge, Schneider, Davidson, Ogshea, Sohal, Goshen, Finkelstein, Paz, Stehfest, Fudim, Ramakrishnan, Huguenard, Hegemann and Deisseroth2011) and ChIEF (Lin et al., Reference Lin, Lin, Steinbach and Tsien2009) exhibit very small desensitizations, of about 10–20% during typical continuous illumination, and thus show consistent reliability at modest light powers (2 mW/mm2) (Mattis et al., Reference Mattis, Tye, Ferenczi, Ramakrishnan, O’Shea, Prakash, Gunaydin, Hyun, Fenno, Gradinaru, Yizhar and Deisseroth2011). Of course, the probability of spike elicitation depends on the overall photocurrent amplitude, and not just the kinetics – thus a depolarizing CCR with a very large photocurrent, even if it has suboptimal kinetics, could still be useful for driving spikes with high probability.
One complicating factor is that all of these emergent properties – I steady-state/I peak, τ desensitization, and the photocurrent magnitude – are generated by a population of rhodopsins, each engaged in a photocycle that is driven by received photons, and once the photocycle has begun, is governed in a stochastic fashion by internal as well as external parameters. Thus, these macroscopically measurable biophysical parameters are affected by the applied light power density and wavelength, which are thus key parameters to take into consideration during opsin selection, especially for in vivo application in mammals, where light absorbance (e.g., by blood) and scattering (e.g., by lipids) mean that different neurons might receive different amounts of light power when light is delivered in typical fashion, for example, from an LED, laser, or optical fiber. Some rules of thumb are useful to consider. For example, in general, at higher light power densities, desensitization rate increases. Thus, rhodopsins with high light sensitivity require lower light power densities and, therefore, will, in general, engage less light-dependent τ desensitization augmentation during a typical neuroscience experiment.
As with photocurrent, light sensitivity can be represented as a single molecule characteristic, or as a cumulative property of the entire set of functional rhodopsin molecules in a given neuron. Single-molecule light sensitivity is an intrinsic opsin property, which is determined by light activation efficiency – a product of extinction coefficient (photon absorption cross-section, perhaps in the vicinity of ~50,000 M−1 cm−1) (Beckmann and Hegemann, Reference Beckmann and Hegemann1991; Muders et al., Reference Muders, Kerruth, Lórenz-Fonfría, Bamann, Heberle and Schlesinger2014)and quantum yield (probability of an opsin advancing to the next stage of the photocycle, upon absorption of one photon, which is in the range from 30 to 80% across rhodopsins) (Ernst et al., Reference Ernst, Lodowski, Elstner, Hegemann, Brown and Kandori2014). However, effective light sensitivity, measured on an ensemble of functional opsin molecules in a cell, is a much more practical way to characterize the light-dependent performance of rhodopsins. To first order, effective light sensitivity can be quantitatively represented by the light intensity required to achieve half-maximal photocurrent, or effective power density for 50% activation (EDP50; Figure 5). High light sensitivity, corresponding to a lower EDP50 value, facilitates stimulation of larger volumes of tissue for a given light power, and reduces phototoxicity on illuminated cells because less light power is required for a given desired volume of illumination. This property also can enable less invasive modulation of cells, or control of neurons far from a light source. For light-gated ion channels, effective light sensitivity exhibits a strong correlation with the rate of channel closure, measured after a light pulse shuts off (Mattis et al., Reference Mattis, Tye, Ferenczi, Ramakrishnan, O’Shea, Prakash, Gunaydin, Hyun, Fenno, Gradinaru, Yizhar and Deisseroth2011). A slower channel closure rate means that more ions are transported into or out of the cell per light pulse, all other things equal – and thus corresponds to higher light sensitivity; thus, there is a tradeoff between off kinetics and light sensitivity. Of course, experiment-dependent conditions like the nature of light propagation in a specific biological system, or varying expression levels of opsin proteins, mean that while the biophysical characteristics here discussed are useful in choosing an opsin for an application, some optimization may be required for any given experiment.
Opsin kinetics affects another neuroscience experiment parameter – the temporal precision of optogenetic control. Photocurrent rise and fall rates contribute to the temporal precision of action potential activation or silencing, upon light pulse delivery to an opsin-expressing neurons. For cation ChRs, on kinetics is often designated by a parameter τ on, defined as time after a light pulse begins to reach peak photocurrent (Berndt et al., Reference Berndt, Schoenenberger, Mattis, Tye, Deisseroth, Hegemann and Oertner2011; Mattis et al., Reference Mattis, Tye, Ferenczi, Ramakrishnan, O’Shea, Prakash, Gunaydin, Hyun, Fenno, Gradinaru, Yizhar and Deisseroth2011) or, in some cases, as time to 90% of the peak (Chater et al., Reference Chater, Henley, Brown and Randall2010; Klapoetke et al., Reference Klapoetke, Murata, Kim, Pulver, Birdsey-Benson, Cho, Morimoto, Chuong, Carpenter, Tian, Wang, Xie, Yan, Zhang, Chow, Surek, Melkonian, Jayaraman, Constantine-Paton, Wong and Boyden2014), when a typically short (2–5 ms) and bright light pulse is delivered, as is common in spike-driving optogenetics experiments (Figure 6a). For ion pumps and anion channelrhodopsins, kinetic parameters are measured under longer light stimulations, typically 1 s or longer. Because overall photocurrent results from the balance between channel opening rate, and desensitization, the time to peak is an emergent property of the applied light power density, and as a result can vary by an order of magnitude from one experimental condition to another, for the same opsin. In general, a higher light power density will result in a shorter time-to-peak.
Another commonly discussed parameter, useful when choosing a depolarizing CCR for a neuroscience experiment, is the temporal precision of the action potentials that result from a light pulse delivery. The time to the peak of an action potential, after light onset, often called the action potential latency of a channelrhodopsin variant, and depends on not only the kinetics of the opsin, but also its photocurrent (Chaigneau et al., Reference Chaigneau, Ronzitti, Gajowa, Soler-Llavina, Tanese, Brureau, Papagiakoumou, Zeng and Emiliani2016; Ronzitti et al., Reference Ronzitti, Conti, Zampini, Klapoetke, Tanese, Papagiakoumou, Boyden, Papagiakoumou and Emiliani2016; Shemesh et al., Reference Shemesh, Tanese, Zampini, Changyang, Kiryln, Ronzitti, Papagiakoumou, Boyden and Emiliani2017). Another parameter describing the temporal precision of light-evoked action potentials is the jitter, defined as the standard deviation of the action potential latency; a high jitter means that spike trains will be noisier and more unreliable in timing. Due to the desensitization of depolarizing CCRs, latency, as well as jitter, generally increase during trains of light flashes (Chater et al., Reference Chater, Henley, Brown and Randall2010). Therefore, an opsin with a large and stable photocurrent can support higher temporal precision action potential trains, as has been seen, for example, with the high conductance depolarizing CCR CoChR (Shemesh et al., Reference Shemesh, Tanese, Zampini, Linghu, Piatkevich, Ronzitti, Papagiakoumou, Boyden and Emiliani2017).
Unlike the rise time, the decay of photocurrent after the termination of a light pulse does not show light intensity dependence. In regard to ChRs, photocurrent generally decays biexponentially following light offset (Figure 6a), whereas for light-driven pumps, the closing kinetics is monoexponential. For convenience, closing rate, abbreviated as the off kinetic parameter τ off, is usually reported as a single value, obtained either from a monoexponential fit of the decay, or as the weighted linear combination of the two-time constants defining a bi-exponential curve. For depolarizing CCRs, fast-off kinetics help with the avoiding of sustained depolarization after a light pulse ends; continued depolarization could result in excess spikes for a given light pulse, with usually uncontrolled timing. When light pulses are delivered at high frequencies, and if each light pulse is desired to result in one precisely timed spike, then fast-off kinetics can prevent continued depolarization from one light pulse from interfering with the depolarization caused by the following light pulse. Stimulation of channelrhodopsins with light pulses delivered faster than the channel-closing rate results in an accumulation of open channelrhodopsin molecules, which can result in an enduring plateau of depolarization – which can lead to uncontrolled spiking, or spike failures if endogenous sodium channels inactivate from the sustained depolarization (Mattis et al., Reference Mattis, Tye, Ferenczi, Ramakrishnan, O’Shea, Prakash, Gunaydin, Hyun, Fenno, Gradinaru, Yizhar and Deisseroth2011; Herman et al., Reference Herman, Huang, Murphey, Garcia and Arenkiel2014).
More generally, since the physiological range of membrane potentials experienced by neurons is quite wide, ranging typically from −80 mV to +10 mV, but often going to even greater extremes, from below −100 mV to beyond +50 mV, it is important to keep in mind that the photocurrent magnitudes of all rhodopsins exhibit voltage dependence. Within a class of opsin, overall I–V trends will generally follow a specific pattern (Figure 6c), and thus I–V curve shape may not be the most critical selection criterion for choosing a specific opsin from a class of phenotypically similar molecules; nevertheless, the I–V curve does represent a fundamental property important for understanding the biophysical principles of optogenetics. The voltage dependence of a photocurrent is characterized by the reversal potential E rev, defined as the membrane potential corresponding to zero photocurrent. I–V curves for depolarizing CCRs are typically asymmetric, except for examples like Chrimson (Vierock et al., Reference Vierock, Grimm, Nitzan and Hegemann2017), showing higher inward conductance at more negative membrane potential values, with a reversal potential close to 0 mV (Feldbauer et al., Reference Feldbauer, Zimmermann, Pintschovius, Spitz, Bamann and Bamberg2009; Chater et al., Reference Chater, Henley, Brown and Randall2010; Gradmann et al., Reference Gradmann, Berndt, Schneider and Hegemann2011). Naturally occurring anion ChRs, as well as newer engineered chloride ChRs, have E rev similar to that of the chloride anion in a given cell, and exhibit linear current–voltage relationships (Govorunova et al., Reference Govorunova, Sineshchekov, Janz, Liu and Spudich2015, Reference Govorunova, Sineshchekov, Rodarte, Janz, Morelle, Melkonian, Gane and Spudich2017; Wietek et al., Reference Wietek, Beltramo, Scanziani, Hegemann, Oertner and Wiegert2015; Berndt et al., Reference Berndt, Lee, Wietek, Ramakrishnan, Steinberg, Rashid, Kim, Park, Santoro, Frankland, Iyer, Pak, Ährlund-Richter, Delp, Malenka, Josselyn, Carlén, Hegemann and Deisseroth2016), unlike those for cation ChRs. In contrast to ChRs, the reversal potential of light-driven ion pumps is extremely negative, because pumps dissipate energy in the service of ion transport and thus can go against a concentration gradient; assuming linear current–voltage relationships beyond the physiological range of membrane potentials, one can extrapolate that the reversal potential may fall in the range of −300 to −400 mV (Seki et al., Reference Seki, Miyauchi, Hayashi, Kikukawa, Kubo, Demura, Ganapathy and Kamo2007; Chow et al., Reference Chow, Han, Dobry, Qian, Chuong, Li, Henninger, Belfort, Lin, Monahan and Boyden2010; Chuong et al., Reference Chuong, Miri, Busskamp, Matthews, Acker, Sørensen, Young, Klapoetke, Henninger, Kodandaramaiah, Ogawa, Ramanlal, Bandler, Allen, Forest, Chow, Han, Lin, Tye, Roska, Cardin and Boyden2014). This property of light-driven pumps means that unnatural distributions of ions can arise from extensive pump use, which in turn could result in artifacts in controlling physiology, as discussed below.
Action spectrum
Action spectrum, the dependence of photocurrent magnitude on illumination wavelength, defines the optimal wavelength for opsin activation and governs how multiple rhodopsins can be used in the same system, or how an opsin can be used in conjunction with simultaneous neural activity imaging. In addition, with red light going deeper in the brain than other colors of visible light, due to lower levels of absorption, seeking red-shifted rhodopsins has been a priority to enable larger volumes of brain tissue to be illuminated, with lower light powers. Maxima of spectral responses for rhodopsins published to date span a wide range of wavelengths, from 435 to 605 nm (Figure 4). In addition, action spectrum shapes can vary a lot, with the full width at half maximum ranging from ~100 to ~200 nm, most likely due to the action spectrum reflecting a superposition of data from multiple chromophore states, possessing different absorption properties. Due to the wide action spectra of rhodopsins, compared with the full width at half maximum of GFP-like fluorescent proteins (30–70 nm), spectrally multiplexed optogenetic control is possible for no more than two rhodopsins, each chosen from an extreme of the spectral palette, and typically requires fine-tuning of light intensities, protein expression level, and stimulation pulse duration, for successful independent control of multiple rhodopsins in the same system (Klapoetke et al., Reference Klapoetke, Murata, Kim, Pulver, Birdsey-Benson, Cho, Morimoto, Chuong, Carpenter, Tian, Wang, Xie, Yan, Zhang, Chow, Surek, Melkonian, Jayaraman, Constantine-Paton, Wong and Boyden2014; Hooks, Reference Hooks2018). Coexpression of blue-driven GtACR2 and red-driven Chrimson enabled sensitive, reliable control of neuronal silencing and spiking, respectively, within the same cell (Vierock et al., Reference Vierock, Rodriguez-Rozada, Dieter, Pieper, Sims, Tenedini, Amelie, Bendifallah, Zhou, Zeitzschel, Ahlbeck, Augustin, Sauter, Papagiakoumou, Gottschalk, Soba, Emiliani, Engel, Hegemann and Wiegert2021). The blue shoulder exhibited by all action spectra also makes it challenging to combine even the furthest red-shifted rhodopsins with optical read-out using common GFP-derived biosensors, because continuous blue light illumination, even at low light powers (e.g., 0.1 mW/mm2, as used for GFP imaging), can be integrated by rhodopsins over extended imaging periods, sometimes causing substantial alternations in membrane potential (Klapoetke et al., Reference Klapoetke, Murata, Kim, Pulver, Birdsey-Benson, Cho, Morimoto, Chuong, Carpenter, Tian, Wang, Xie, Yan, Zhang, Chow, Surek, Melkonian, Jayaraman, Constantine-Paton, Wong and Boyden2014; Trojanowski et al., Reference Trojanowski, Nelson, Flavell, Fang-Yen and Raizen2015). Using red and near-infrared sensors in conjunction with blue-light activated rhodopsins avoids this issue, and may yield a more straightforward approach for spectral multiplexing of neural control and imaging (Piatkevich et al., Reference Piatkevich, Jung, Straub, Linghu, Park, Suk, Hochbaum, Goodwin, Pnevmatikakis, Pak, Kawashima, Yang, Rhoades, Shemesh, Asano, Yoon, Freifeld, Saulnier, Riegler, Engert, Hughes, Drobizhev, Szabo, Ahrens, Flavell, Sabatini and Boyden2018; Piatkevich et al., Reference Piatkevich, Murdock and Subach2019; Qian et al., Reference Qian, Piatkevich, Larney, Abdelfattah, Mehta, Murdock, Gottschalk, Molina, Zhang, Chen, Wu, Drobizhev, Hughes, Zhang, Schreiter, Shoham, Razansky, Boyden and Campbell2019, Reference Qian, Cosio, Piatkevich, Aufmkolk, Su, Celiker, Schohl, Murdock, Aggarwal, Chang, Wiseman, Ruthazer, Boyden and Campbell2020). However, to date, there are relatively few red-shifted sensors of neuronal activity available (Lin and Schnitzer, Reference Lin and Schnitzer2016; Piatkevich et al., Reference Piatkevich, Bensussen, Tseng, Shroff, Lopez-Huerta, Park, Jung, Shemesh, Straub, Gritton, Romano, Costa, Sabatini, Fu, Boyden and Han2019).
Although fundamentally and technically more challenging than wide-field one-photon illumination, two-photon activation of individual neurons offers excellent multiplexing capability, because light can be directed to a targeted cell and not others, with high spatial resolution, even in scattering tissue such as in the living mammalian brain (Oron et al., Reference Oron, Eirini Papagiakoumou and Emiliani2012; Papagiakoumou et al., Reference Papagiakoumou, Ronzitti and Emiliani2020). The two-photon action spectrum is generally not fully predictable, considering just the one-photon action spectrum; however, the two-photon action spectrum maximum is approximately two times that of the one-photon action spectrum maximum. A standard Ti-Sapphire laser, as widely used in two-photon microscopy, was sufficient for both in vitro and in vivo photostimulation of multiple ChRs with single neuron spatial precision (Rickgauer and Tank, Reference Rickgauer and Tank2009; Andrasfalvy et al., Reference Andrasfalvy, Zemelman, Tang and Vaziri2010; Packer et al., Reference Packer, Russell, Henry and Häusser2015; Shemesh et al., Reference Shemesh, Tanese, Zampini, Linghu, Piatkevich, Ronzitti, Papagiakoumou, Boyden and Emiliani2017), as well as engagement of ion pumps (Carrillo-Reid et al., Reference Carrillo-Reid, Han, Yang, Akrouh and Yuste2019; Chen et al., Reference Chen, Ronzitti, Lee, Daigle, Dalkara, Zeng, Emiliani and Papagiakoumou2019; Marshel et al., Reference Marshel, Kim, Machado, Quirin, Benson, Kadmon, Raja, Chibukhchyan, Ramakrishnan, Inoue, Shane, DJ, Yoshizawa, Kato, Ganguli and Deisseroth2019). Recent advances in optical illumination methods enable simultaneous two-photon photostimulation of many neurons within a volume of interest, with single-cell (or even subcellular) resolution, with millisecond timescale precision (Pégard et al., Reference Pégard, Mardinly, Oldenburg, Sridharan, Waller and Adesnik2017; Shemesh et al., Reference Shemesh, Tanese, Zampini, Linghu, Piatkevich, Ronzitti, Papagiakoumou, Boyden and Emiliani2017; Mardinly et al., Reference Mardinly, Oldenburg, Pégard, Sridharan, Lyall, Chesnov, Brohawn, Waller and Adesnik2018) in a fashion that can be combined with two-photon imaging of targeted cells (Peterka et al., Reference Peterka, Takahashi and Yuste2011; Carrillo-Reid et al., Reference Carrillo-Reid, Han, Yang, Akrouh and Yuste2019; Marshel et al., Reference Marshel, Kim, Machado, Quirin, Benson, Kadmon, Raja, Chibukhchyan, Ramakrishnan, Inoue, Shane, DJ, Yoshizawa, Kato, Ganguli and Deisseroth2019).
New high-power laser setups are beginning to enable three-photon optogenetics, with the potential for deeper penetration into the brain, although this has been only demonstrated in cultured neurons so far (Rowlands et al., Reference Rowlands, Park, Bruns, Piatkevich, Fukumura, Jain, Bawendi, Boyden and So2016). In general, the properties of the photon-excited state of a rhodopsin do not depend on the way it was excited, so the fundamental biophysical properties of rhodopsins associated with the excited state under two-photon illumination, such as I steady-state/I peak, τ desensitization, τ recovery, and τ off, should correspond to those measured under one-photon activation. However, under two-photon excitation, in which light arrives in subpicosecond duration pulses at high (e.g., 500 kHz–80 MHz) repetition rates, rather than the continuous flux of photons seen in one-photon optogenetics, the photocurrent achievable, and the τ on, may depend on the details of the illumination used, including properties of the laser. Since photoactivation is typically spatially multiplexed with two-photon control, for example, a scanning laser might have to be steered to different neurons at different points of time; since photoactivation occurs only during illumination periods, slow channel off-kinetics can be beneficial for accumulating open channelrhodopsins over illumination periods, eventually contributing to higher maximum photocurrent (Packer et al., Reference Packer, Peterka, Hirtz, Prakash, Deisseroth and Yuste2012; Prakash et al., Reference Prakash, Yizhar, Grewe, Ramakrishnan, Wang, Goshen, Packer, Peterka, Yuste, Schnitzer and Deisseroth2012). Nevertheless, two-photon activation of the ultrafast depolarizing ChR Chronos (Klapoetke et al., Reference Klapoetke, Murata, Kim, Pulver, Birdsey-Benson, Cho, Morimoto, Chuong, Carpenter, Tian, Wang, Xie, Yan, Zhang, Chow, Surek, Melkonian, Jayaraman, Constantine-Paton, Wong and Boyden2014) and of a proton pump with fast photocycle, Arch (Chow et al., Reference Chow, Han, Dobry, Qian, Chuong, Li, Henninger, Belfort, Lin, Monahan and Boyden2010), has been shown to be sufficient for optogenetic control of neurons in acute brain slice (Prakash et al., Reference Prakash, Yizhar, Grewe, Ramakrishnan, Wang, Goshen, Packer, Peterka, Yuste, Schnitzer and Deisseroth2012; Ronzitti et al., Reference Ronzitti, Conti, Zampini, Klapoetke, Tanese, Papagiakoumou, Boyden, Papagiakoumou and Emiliani2016).
Ion selectivity
Optogenetic tools transport specific ions, which may exist in different concentrations in different neural states, and which can have different effects on downstream physiology. Thus, it is important to consider the ion selectivity of a given rhodopsin, to understand and to be able to predict the impact of the usage of a given opsin on the biochemical processes of a cell. Rhodopsins exhibiting a diversity of ion selectivities have been discovered and characterized. Some rhodopsins have been engineered for altered ion selectivity, thus expanding the neuroscientist’s toolbox. We have discussed, throughout this paper, four major classes of rhodopsins of wide application in neuroscience – cation-conducting channelrhodopsins (also known as just channelrhodopsin, ChRs, or CCRs as used by some authors), which are typically depolarizing except for the recently discovered class of K+-conducting channelrhodopsins (Govorunova et al., Reference Govorunova, Gou, Sineshchekov, Li, Lu, Wang, Brown, St-Pierre, Xue and Spudich2022; Vierock et al. Reference Vierock, Peter, Grimm, Rozenberg, Scalise, Augustin, Tanese, Forget, Emiliani, Béjà and Hegemann2022); anion-conducting channelrhodopsins (also referred to as ACRs); inward light-driven chloride pumps; and outward light-driven proton pumps. For each class of rhodopsins, the name indicates the types of ions each rhodopsin is selective for, and CCRs are additionally subcategorized as depolarizing or hyperpolarizing; colloquially, the word ‘cation’ is sometimes dropped from the phrase cation ChR, since the first ChRs to be used in neuroscience were all cation-conducting and thus sometimes ChR, when used alone, refers to a depolarizing cation ChR2 (Boyden et al., Reference Boyden, Zhang, Bamberg, Nagel and Deisseroth2005). ACRs derived from ChRs via protein engineering are sometimes referred to as designed or engineered ACRs (dACRs (Kato et al., Reference Kato, Kim, Paggi, Evans, Allen, Richardson, Inoue, Ito, Ramakrishnan, Fenno, Yamashita, Hilger, Lee, Berndt, Shen, Kandori, Dror, Kobilka and Deisseroth2018) or eACRs (Wietek et al., Reference Wietek, Rodriguez-Rozada, Tutas, Tenedini, Grimm, Oertner, Soba, Hegemann and Wiegert2017), respectively, for short).
All studied cation ChRs conduct protons and physiologically relevant monovalent and bivalent metal cations, such as sodium, potassium, calcium, and magnesium, with inward rectification (note the asymmetric I–V curves in Figure 6c). All ChRs are ion-selective, with the following relative conductivities typical: H+ > > Na+ > K+ > Ca2+ > Mg2+, likely because of the differential binding affinity of ions to key amino acid residues within the pore (Schneider et al., Reference Schneider, Gradmann and Hegemann2013). Relative ion conductivities vary across ChR species, and show strong voltage- and pH-dependence (Schneider et al., Reference Schneider, Gradmann and Hegemann2013), meaning that the ion composition of the photocurrent depends on the existing ion gradients across the plasma membrane. For example, for ChR2 at more negative membrane voltages, the sodium photocurrent is several times higher than when measured at a voltage closer to the reversal potential of sodium, where photocurrent is more carried by protons (Berndt et al., Reference Berndt, Prigge, Gradmann and Hegemann2010; Schneider et al., Reference Schneider, Gradmann and Hegemann2013). For ChR2, despite its very high selectivity toward protons (the selectivity ratio, PH+/PNa+, is about 2 × 106) (Nagel et al., Reference Nagel, Szellas, Huhn, Kateriya, Adeishvili, Berthold, Ollig, Hegemann and Bamberg2003; Berndt et al., Reference Berndt, Prigge, Gradmann and Hegemann2010; Vierock et al., Reference Vierock, Grimm, Nitzan and Hegemann2017), under physiological conditions common in the brain, where the extracellular concentration of sodium is ~150 mM and the pH is ~7.3–7.4, about half of the photocurrent is carried by protons. The rest of the photocurrent is carried mainly by sodium, with a small fraction of calcium and magnesium ions, while the contribution of potassium current is negligible due to its higher concentration inside cells (and such ChRs are inwardly rectifying).
Structure–function relationships for ion selectivity in rhodopsin are still poorly understood, and therefore the rational design of an entirely ion-selective cation channel has been very challenging. There exist multiple engineered and naturally occurred ChR variants with improved sodium, calcium, or magnesium conductance; however, their improved cation selective properties arose as much, or more, from serendipity than from rational design. For example, the first generated point mutant of ChR2, the H134R mutant, sometimes called ChR2R, has found widespread application in neuroscience and exhibits a modest increase in sodium current compared to its precursor (Gradmann et al., Reference Gradmann, Berndt, Schneider and Hegemann2011). Another wild-type channelrhodopsin, PsChR from Platymonas tetraselmis subcordiformis, has one of the highest sodium selectivities among all studied wild-type channelrhodopsin (PH+/PNa+ ~ 5 ·105) (Govorunova et al., Reference Govorunova, Sineshchekov, Li, Janz and Spudich2013; Duan et al., Reference Duan, Nagel and Gao2019). The D139H mutation of PsChR further increased Na+ selectivity, over H+, by fivefold. Furthermore, PsChR D139H showed a fivefold larger photocurrent than wild-type PsChR. Interestingly, the single amino acid substitution E143S, in the ion pore of Chrimson (called ChrimsonS), increased sodium selectivity by more than two orders of magnitude, with PH+/PNa+ going from 1.3 × 107 to 5.3 × 105, thus significantly altering the ion composition of the photocurrent. To put this into context: under physiological conditions, 90% of Chrimson’s photocurrent is carried by protons; however, in the case of ChrimsonS, only 20% of the photocurrent consists of protons. This increase in selectivity, however, comes at the cost of reduced photocurrent – by about 2.5-fold. This is one of the reasons opsin engineering is challenging – it is hard to change one property of an opsin completely independently of all other properties of an opsin. In addition, red-shifted channelrhodopsins, such as the C1V1 chimera and its accelerated variants, have increased conductance for calcium and magnesium (Prigge et al., Reference Prigge, Schneider, Tsunoda, Shilyansky, Wietek, Deisseroth and Hegemann2012; Schneider et al., Reference Schneider, Gradmann and Hegemann2013), although another channelrhodopsin with very high calcium conductance, named calcium translocating channelrhodopsin (CatCh) and its improved variant CatCh+, are mutants of ChR2 (Li et al., Reference Li, Karine, Isacoff, Oheim and Ropert2012; Prigge et al., Reference Prigge, Schneider, Tsunoda, Shilyansky, Wietek, Deisseroth and Hegemann2012; Kim et al. Reference Kim, Rouault, Druckmann and Jayaraman2017; Mager et al., Reference Mager, Wood and Bamberg2017).
In terms of the wild-type ChR2, significant conductance of calcium ions occurs only under certain conditions, such as high extracellular calcium concentration, created artificially (Caldwell et al., Reference Caldwell, Herin, Nagel, Bamberg, Scheschonka and Betz2008; Schneider et al., Reference Schneider, Gradmann and Hegemann2013), or under high local intracellular calcium concentration, for example, originating from intracellular Ca stores (Figueiredo et al., Reference Figueiredo, Lane, Stout, Liu, Parpura, Teschemacher and Kasparov2014). It is common to observe an elevation in intracellular [Ca2+] upon ChR2 photoactivation in neurons, but this is likely often due primarily to the secondary activation of voltage-gated calcium channels by ChR2-mediated depolarization (Zhang and Oertner, Reference Zhang and Oertner2007; Li et al., Reference Li, Karine, Isacoff, Oheim and Ropert2012). Therefore, the interpretation of any observed changes in ion concentration, upon optogenetic stimulation, must take into account the endogenous channels and pumps responsible for neural function.
Of course, whether an ion channel or pump results in a depolarizing or hyperpolarizing effect depends on the details of the cell’s physiology. As an example, the chloride gradient across the plasma membrane can differ in neurons at various developmental stages (Kaila et al., Reference Kaila, Price, Payne, Puskarjov and Voipio2014; Heigele et al., Reference Heigele, Sultan, Toni and Bischofberger2016; Raimondo et al., Reference Raimondo, Richards and Woodin2017; Sato et al., Reference Sato, Artoni, Landi, Parra, Trovato, Szczurkowska, Luin, Arosio, Beltram, Cancedda, Kaila and Ratto2017), across neuronal compartments (Turecek and Trussell, Reference Turecek and Trussell2001; Price and Trussell, Reference Price and Trussell2006; Szabadics et al., Reference Szabadics, Varga, Molnar, Olah, Barzo and Tamas2006; Khirug et al., Reference Khirug, Yamada, Afzalov, Voipio, Khiroug and Kaila2008; Pugh and Jahr, Reference Pugh and Jahr2011) and across normal versus pathological conditions (Cohen et al., Reference Cohen, Navarro, Clemenceau, Baulac and Miles2002; Huberfeld et al., Reference Huberfeld, Wittner, Clemenceau, Baulac, Kaila, Miles and Rivera2007; Price et al., Reference Price, Cervero, Gold, Hammond and Prescott2009; Boulenguez et al., Reference Boulenguez, Liabeuf, Bos, Bras, Jean-Xavier, Brocard, Stil, Darbon, Cattaert, Delpire, Marsala and Vinay2010; Tao et al., Reference Tao, Li, Newburn, Ye, Lipska, Herman, Weinberger, Kleinman and Hyde2012; Nelson and Valakh, Reference Nelson and Valakh2015; Tang et al., Reference Tang, Kim, Zhou, Wengert, Zhang, Wu, Carromeu, Muotri, Marchetto, Gage and Chen2016). For example, under normal conditions the cytoplasmic [Cl–] ~ 4–7 mM in somata of mature neurons (Bregestovski et al., Reference Bregestovski, Waseem and Mukhtarov2009; Sato et al., Reference Sato, Artoni, Landi, Parra, Trovato, Szczurkowska, Luin, Arosio, Beltram, Cancedda, Kaila and Ratto2017) is lower than extracellular [Cl–], and thus activation of anion channelrhodopsins results in inward-directed photocurrent, shunting depolarization of the cell to the reversal potential of chloride, which is usually near the resting membrane potential (Wietek et al., Reference Wietek, Beltramo, Scanziani, Hegemann, Oertner and Wiegert2015; Berndt et al., Reference Berndt, Lee, Wietek, Ramakrishnan, Steinberg, Rashid, Kim, Park, Santoro, Frankland, Iyer, Pak, Ährlund-Richter, Delp, Malenka, Josselyn, Carlén, Hegemann and Deisseroth2016; Chung et al., Reference Chung, Weber, Zhong, Tan, Nguyen, Beier, Hörmann, Chang, Zhang, Do, Yao, Krashes, Tasic, Cetin, Zeng, Knight, Luo and Dan2017; Zhang et al., Reference Zhang, Zhou, Chen, Ni, Zhao, Yang, Gao, Fu, Zhang and Zhou2017). Axons can accumulate three to five times higher [Cl–] than in their parent cell bodies, however (Price and Trussell, Reference Price and Trussell2006; Khirug et al., Reference Khirug, Yamada, Afzalov, Voipio, Khiroug and Kaila2008), so that even brief illumination (10 ms) of axons expressing GtACRs, chloride specific channelrhodopsins, could cause presynaptic release (Mahn et al., Reference Mahn, Prigge, Ron, Levy and Yizhar2016) and evoke antidromic spikes (Malyshev et al., Reference Malyshev, Roshchin, Smirnova, Dolgikh, Balaban and Ostrovsky2017) in acute brain slices, due to outward chloride photocurrent resulting in light-driven depolarization. It should be noted that selective illumination of somata of the same neuron types efficiently inhibited action potentials (Malyshev et al., Reference Malyshev, Roshchin, Smirnova, Dolgikh, Balaban and Ostrovsky2017).
In contrast, light-driven chloride pumps can hyperpolarize neurons across a wide range of conditions, due to the active transport of chloride ions into cells under illumination, which is largely of the chloride gradient or the membrane potential across the membrane (Han and Boyden, Reference Han and Boyden2007; Gradinaru et al., Reference Gradinaru, Thompson and Deisseroth2008; Mattis et al., Reference Mattis, Tye, Ferenczi, Ramakrishnan, O’Shea, Prakash, Gunaydin, Hyun, Fenno, Gradinaru, Yizhar and Deisseroth2011; Chuong et al., Reference Chuong, Miri, Busskamp, Matthews, Acker, Sørensen, Young, Klapoetke, Henninger, Kodandaramaiah, Ogawa, Ramanlal, Bandler, Allen, Forest, Chow, Han, Lin, Tye, Roska, Cardin and Boyden2014). However, even brief activation of chloride pumps in neurons (1–10s) leads to an increase in intracellular chloride concentration, which can cause positive shifts in the GABAergic reversal potential (Raimondo et al., Reference Raimondo, Kay, Ellender and Akerman2012; Alfonsa et al., Reference Alfonsa, Merricks, Codadu, Cunningham, Deisseroth, Racca and Trevelyan2015), which can induce rebound activity. Rebound activity also can be triggered by hyperpolarization-activated Ih currents (Biel et al., Reference Biel, Wahl-Schott, Michalakis and Zong2009; Tonnesen et al., Reference Tonnesen, Sorensen, Deisseroth, Lundberg and Kokaia2009). Thus, following illumination to photoinhibit cells, increased firing rates have been observed both in acute slice preparations (Raimondo et al., Reference Raimondo, Kay, Ellender and Akerman2012; Alfonsa et al., Reference Alfonsa, Merricks, Codadu, Cunningham, Deisseroth, Racca and Trevelyan2015) and in vivo in mice (Madisen et al., Reference Madisen, Mao, Koch, Zhuo, Berenyi, Fujisawa, Hsu, Garcia, Gu, Zanella, Kidney, Gu, Mao, Hooks, Boyden, Buzsáki, Ramirez, Jones, Svoboda, Han, Turner and Zeng2012; Chuong et al., Reference Chuong, Miri, Busskamp, Matthews, Acker, Sørensen, Young, Klapoetke, Henninger, Kodandaramaiah, Ogawa, Ramanlal, Bandler, Allen, Forest, Chow, Han, Lin, Tye, Roska, Cardin and Boyden2014) and zebrafish (Arrenberg et al., Reference Arrenberg, Bene and Baier2009), thus making it important to characterize how photoactivation of a rhodopsin will change the voltage or firing activity of a particular cell type when using these tools. Similarly, illumination of light-driven proton pumps for extended periods of time could increase spontaneous presynaptic transmitter release, perhaps by facilitating a calcium influx (Mahn et al., Reference Mahn, Prigge, Ron, Levy and Yizhar2016).
Although changes in cellular pH driven by light-driven proton pumps are numerically small (e.g., 0.1–0.2 pH units for a typical illumination pattern in a neuron (Chow et al., Reference Chow, Han, Dobry, Qian, Chuong, Li, Henninger, Belfort, Lin, Monahan and Boyden2010)), local changes, potentially coupled to the presence of specific pH-sensitive proteins in certain cell types or compartments, may respond to such changes. In short, neural silencing must be carefully thought through, because the long durations over which optogenetic silencers are typically utilized, mean that changes in ion concentrations must be considered, and controlled for. The recent discovery of light-driven potassium channels may offer an alternative to the above reagents, by helping avoid artifacts associated with chloride or protons (Govorunova et al., Reference Govorunova, Gou, Sineshchekov, Li, Lu, Wang, Brown, St-Pierre, Xue and Spudich2022; Vierock et al. Reference Vierock, Peter, Grimm, Andrey Rozenberg, Tillert, Scalise, Casini, Augustin, Tanese, Forget, Peyronnet, Schneider-Warme, Emiliani, Béjà and Hegemann2022) And, other pumps are being discovered, which may have uses in neuroscience. For example, a light-driven sodium pump, KR2, was discovered in Krokinobacter eikastus (Inoue et al., Reference Inoue, Ono, Abe-Yoshizumi, Yoshizawa, Ito, Kogure and Kandori2013). It has potential as a neural silencer, and in a trafficking-enhanced form which boosted membrane expression and photocurrents, denoted eKR2, showed the ability to reduce firing in stimulated cultured neurons in response to 540 nm light of few mW/mm2 irradiance, although to our knowledge it has not been utilized in vivo (Grimm et al., Reference Grimm, Silapetere, Vogt, Sierra and Hegemann2018). Strategically mutagenizing light-driven sodium pumps can impart potassium pumping capability (Gushchin et al., Reference Gushchin, Shevchenko, Polovinkin, Kovalev, Alekseev, Round, Borshchevskiy, Balandin, Popov, Gensch, Fahlke, Bamann, Willbold, Büldt, Bamberg and Gordeliy2015; Kato et al., Reference Kato, Inoue, Abe-Yoshizumi, Kato, Ono, Konno, Hososhima, Ishizuka, Hoque, Kunitomo, Ito, Yoshizawa, Yamashita, Takemoto, Nishizawa, Taniguchi, Kogure, Maturana, Iino, Yawo, Ishitani, Kandori and Nureki2015), opening up yet another potential future direction.
Conclusion
In summary, while some of the properties of rhodopsins that helped them meet Crick’s criteria for success were out-of-the-blue serendipitous – who would have known, for example, that mammalian neurons spontaneously had enough all-trans-retinal around, to enable opsin proteins to function without chemical supplementation? – some of Crick’s criteria were met because of well-understood biophysical properties of rhodopsins. The high speed of operation of rhodopsins arises because of specific properties of their structures, which lend themselves to closed photocycles, favorable kinetics on par with the high speed of neurons, and light sensitivities and action spectra which are well matched to light penetration properties of mammalian brain. The clear mechanisms of action of rhodopsins means that the interpretation of experiments is straightforward, and the presence of alternative choices for some opsin categories (e.g., the neural silencers discussed above) opens up the possibility of experiment-specific customization of reagent use, so that undesired artifacts can be avoided.
Going forward: even as existing opsin toolsets have become widespread in neuroscience, there is much opportunity going forward to apply rhodopsins in even more scientific, and perhaps medical contexts (Sahel et al., Reference Sahel, Boulanger-Scemama, Pagot, Arleo, Galluppi, Martel, Esposti, Delaux, de Saint Aubert, de Montleau, Gutman, Audo, Duebel, Picaud, Dalkara, Blouin, Taiel and Roska2021); new strategies, such as machine learning-assisted directed evolution and-software protein structure prediction may help with further optimization of opsin reagents (Bedbrook et al., Reference Bedbrook, Yang, Robinson, Mackey, Gradinaru and Arnold2019; Jumper et al., Reference Jumper, Evans, Pritzel, Green, Figurnov, Ronneberger, Tunyasuvunakool, Bates, Žídek, Potapenko, Bridgland, Meyer, SAA, Ballard, Cowie, Romera-Paredes, Nikolov, Jain, Adler, Back, Petersen, Reiman, Clancy, Zielinski, Steinegger, Pacholska, Berghammer, Bodenstein, Silver, Vinyals, Senior, Kavukcuoglu, Kohli and Hassabis2021); and synergistic tools such as neural imaging and closed-loop optogenetic control will enable rhodopsins to be used in more and more complex neuroscience question contexts. In some ways the first chapters of the optogenetics story are complete, but in other ways, the adventure is just beginning.
Acknowledgments
We thank Shuguang Zhang for comments on this manuscript. We also thank Siranush Babakhanova and Cuixin Lai for helping with preparing images for Figure 2.
Financial support
This research received no specific grant from any funding agency, commercial, or not-for-profit sectors.
Competing interest
E.S.B. is an inventor on multiple patents related to optogenetics. K.D.P. is an inventor on a patent related to optogenetics.








