Nintedanib (free base CAS Registry Number 656247-17-5) is a drug for the treatment of idiopathic pulmonary fibrosis (IPF) as well as non-small cell lung cancer (NSCLC). In NSCLC, Nintedanib (brand name Vargatef) inhibits three types of growth factor receptors that are involved in angiogenesis for tumor growth as an inhibitor of tyrosine kinases (Yan et al., Reference Yan, Xue, Wang, Gao, Zeng, Wang and Jia2023). In IPF, Nintedanib (brand name Ofev) has antifibrotic properties involving growth factors in the lungs that act against connective tissue formation by inhibiting fibroblast proliferation, which acts similarly to its tumor inhibition properties (Wollin et al., Reference Wollin, Wex, Pautsch, Schnapp, Hostettler, Stowasser and Kolb2015; Marijic et al., Reference Marijic, Schwarzkopf, Schwettmann, Ruhnke, Trudzinski and Kreuter2021). Nintedanib is FDA-approved for Ofev, as an IPF treatment, that has been proven to reduce significant cell proliferation in the lungs against IPF and has been recognized to work well against NSCLC and systemic sclerosis by delaying long-term proliferation. Hepatotoxicity due to increased liver enzymes in metabolism is the main drawback of this type of therapy but has otherwise had promising results for IPF and NSCLC (Gole and Bankole, Reference Gole and Bankole2023).
Primarily administered in a salt form as nintedanib esylate (Figure 1 inset, CAS Registry Number 656247-18-6), the systematic name is methyl (3Z)-3-(((4-(N-methyl-2-(4-methylpiperazin-1-yl)acetamido)phenyl)amino) (phenyl)methylidene)-2-oxo-2,3-dihydro-1H-indole-6-carboxylate ethanesulfonate. This work was carried out as part of a project to determine the crystal structures of large-volume commercial pharmaceuticals and include high-quality powder diffraction data for them in the Powder Diffraction File (Gates-Rector and Blanton, Reference Gates-Rector and Blanton2019).

Figure 1. Nintedanib esylate hemihydrate synchrotron X-ray diffraction pattern. Inset nintedanib esylate hemihydrate molecule. Image generated using JADE Pro (MDI, 2024) and Mercury (Macrae et al., Reference Macrae, Sovago, Cottrell, Galek, McCabe, Pidcock, Platings, Shields, Stevens, Towler and Wood2020).
The synchrotron pattern (Figure 1) of a commercial reagent (purchased from TargetMol, lot #131360) was measured at 11-BM at APS (Antao et al., Reference Antao, Hassan, Wang, Lee and Toby2008; Lee et al., Reference Lee, Shu, Ramanathan, Preissner, Wang, Beno, Von Dreele, Ribaud, Kurtz, Antao, Jiao and Toby2008; Wang et al., Reference Wang, Toby, Lee, Ribaud, Antao, Kurtz, Ramanathan, Von Dreele and Beno2008) using a wavelength of 0.459744 Å). The pattern was indexed using N-TREOR (Altomare et al., Reference Altomare, Cuocci, Giacovazzo, Moliterni, Rizzi, Corriero and Falcicchio2013) on a primitive triclinic unit cell. Triclinic lattice parameters for nintedanib esylate hemihydrate, CAS Registry Number 959762-24-4 (space group P-1, which had a similar unit cell) and a powder pattern were reported by Roth et al. (Reference Roth, Linz, Sieger, Rall, Hilberg and Bock2006; Boehringer Ingelheim Pharma GmbH), but no atom coordinates were included. A connectivity search in the Cambridge Structural Database (Groom et al., Reference Groom, Bruno, Lightfoot and Ward2016) yielded the Refcode YASMIS for nintedanib esylate hemihydrate (Ma et al., Reference Ma, Huang, Cao, Sha, Sun, He, Wan, Li, Li and Ren2022). Although purchased as anhydrous, the sample seems to be the hemihydrate. The sample contains an unknown crystalline impurity. Thirteen peaks from this phase could be indexed on a primitive monoclinic unit cell having a = 14.2907, b = 4.6233, c = 23.6130 Å, β = 91.779°, and V = 1559.37 Å3; this impurity phase was included in the refinement as a Le Bail phase. The structure of the nintedanib esylate hemihydrate phase was refined using GSAS-II (Toby and Von Dreele, Reference Toby and Von Dreele2013) using an ordered model (removing a minor orientation of one of the esylate anions) and optimized using VASP (Kresse and Furthmüller, Reference Kresse and Furthmüller1996). A single-point density functional calculation (fixed experimental cell) and population analysis were carried out using CRYSTAL23 (Erba et al., Reference Erba, Desmaris, Casassa, Civalleri, Donà, Bush, Searle, Maschio, Daga, Cossard, Ribaldone, Ascrizzi, Marana, Flament and Kirtman2022).
The crystal structure contains corrugated layers parallel to the bc-plane. Our lattice parameters (Table I) fall between those of the other two determinations; perhaps, the water content varies among samples. Hydrogen bonds (Table II) are prominent in the crystal structure. The water molecule O13 forms two medium-strength O–H⋯O hydrogen bonds to one of the esylate anions. The protonated nitrogen atom in each cation forms an N–H⋯O hydrogen bond to an esylate anion. The ring N–H groups N31 and N102 form strong intramolecular N–H⋯O hydrogen bonds to carbonyl groups. The ring N–H groups N24 and N104 form intramolecular N–H⋯O hydrogen bonds to esylate anions. Many C–H⋯O hydrogen bonds (and one C–H⋯N hydrogen bond), with aromatic C–H, methylene groups, and methyl groups as donors are present. The hydrogen bonding patterns of the two cations differ considerably. A Mogul geometry analysis indicates that torsion angles involving rotation about the C43–N31 bond are unusual; these fall in a valley in a very broad distribution, so, perhaps, the intramolecular hydrogen bond has resulted in an unusual conformation. The C119–C115 torsion angles are also flagged as unusual.
TABLE I. Reduced lattice parameters of nintedanib esylate hemihydrate. Space group P-1
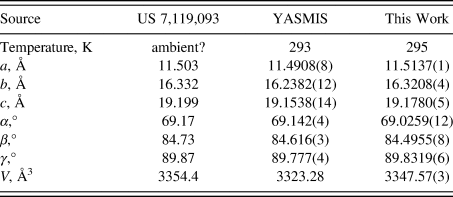
TABLE II. Hydrogen bonds (CRYSTAL23) in nintedanib esylate hemihydrate

a Intramolecular.
I. DEPOSITED DATA
The powder pattern of nintedanib esylate hemihydrate from this synchrotron data set has been submitted to ICDD for inclusion in the Powder Diffraction File. The Crystallographic Information Framework (CIF) files containing the results of the Rietveld refinement (including the raw data) and the DFT geometry optimization were deposited with the ICDD. The data can be requested at [email protected].
ACKNOWLEDGEMENTS
The use of the Advanced Photon Source at Argonne National Laboratory was supported by the U. S. Department of Energy, Office of Science, Office of Basic Energy Sciences, under Contract No. DE-AC02-06CH11357. This work was partially supported by the International Centre for Diffraction Data. We thank Saul Lapidus for his assistance in the data collection.
CONFLICTS OF INTEREST
The authors have no conflicts of interest to declare.






