Introduction
Globally, an estimated 257 million individuals are infected with chronic hepatitis B virus (HBV), which contributed to about 66% of total hepatitis deaths in 2015 (Refs 1, 2). Chronic hepatitis B (CHB) infection is an incurable disease, long-term antiviral treatment is required to suppress viral replication, and disease recurrence is common after treatment discontinuation (Refs Reference Papatheodoridis3–6). Approximately 3% of CHB patients develop liver cirrhosis within one year (Ref. Reference Guan and Lui7) and they have a higher risk of developing hepatocellular carcinoma (HCC) than non-cirrhotic patients (Refs Reference An8–Reference Seto11). Early HCC detection reduces mortality by 37% and increases survival rate of HCC patients by up to 68% with a 5-year follow-up screening (Refs Reference Heimbach12, Reference Zhang, Yang and Tang13). However, this is often not possible because typical symptoms of HCC are not apparent until the disease is advanced (Ref. Reference Zhao, Ju and Li14). Several HCC biomarkers, such as alpha-fetoprotein (AFP), alpha-fetoprotein lens culinaris agglutinin (AFP-L3) and des-gamma carboxy prothrombin (DCP), are used to diagnose HCC, albeit their discrepant diagnostic sensitivity and specificity (Refs Reference Beudeker and Boonstra15–Reference Parikh17).
DNA methylation occurs in the HBV genome as well as in the human genome during CHB infection and hepatocarcinogenesis (Refs Reference Charostad18–Reference Zhang24). Following HBV infection, hepatitis B X (HBx) protein upregulates the activity of DNA methyltransferases (DNMTs) in infected hepatocytes, inducing methylation of viral and host DNA (Refs Reference Lee25, Reference Park26). DNA methylation is an important epigenetic modification that associates with the gene expression (Ref. Reference Lyko27). Several lines of evidence support the role of human tumour suppressor genes (TSGs) methylation in HCC progression (Refs Reference Dong19, Reference Ye23). Methylation of HBV DNA, such as covalently closed circular DNA (cccDNA), has been observed particularly in HCC development (Refs Reference Fernandez20–Reference Kaur22). These findings provide opportunities for HBV and human genome methylation in the diagnosis and prognosis of HBV-related HCC. Here, we review the significance of DNA methylation in the progression of HCC, as well as that of HBV DNA replication and gene expression.
HBV life cycle
HBV is a member of the Hepadnaviridae family, a small DNA virus that has characteristics similar to the retrovirus. It is an enveloped virus with a nucleocapsid. The HBV virion is 42–47 nm in diameter and contains partial double-stranded relaxed circular DNA (rcDNA) with one complete minus strand and one incomplete plus strand. The genome size of HBV is approximately 3.2 kb (Ref. Reference Caballero28). The HBV genome encodes four overlapping open reading frames (ORFs), namely core (C), polymerase (P), surface (S) and X genes (Fig. 1). The C ORF encodes the core and precore proteins. The HBV core (HBc) protein acts as a subunit of the viral nucleocapsid, hepatitis precore (HBe) protein itself will undergo proteolytic process at its N and C termini. P ORF encodes the polyprotein which carries out the synthesis of the viral DNA in its life cycle. The S gene is divided into the pre-S1, pre-S2 and S regions, it encodes three types of viral envelope proteins. X gene plays an important role in viral replication and cellular signal-transduction pathways including cell proliferation and apoptosis. It encodes 16.5-kDa HBx protein (Ref. Reference Seeger and Mason29).
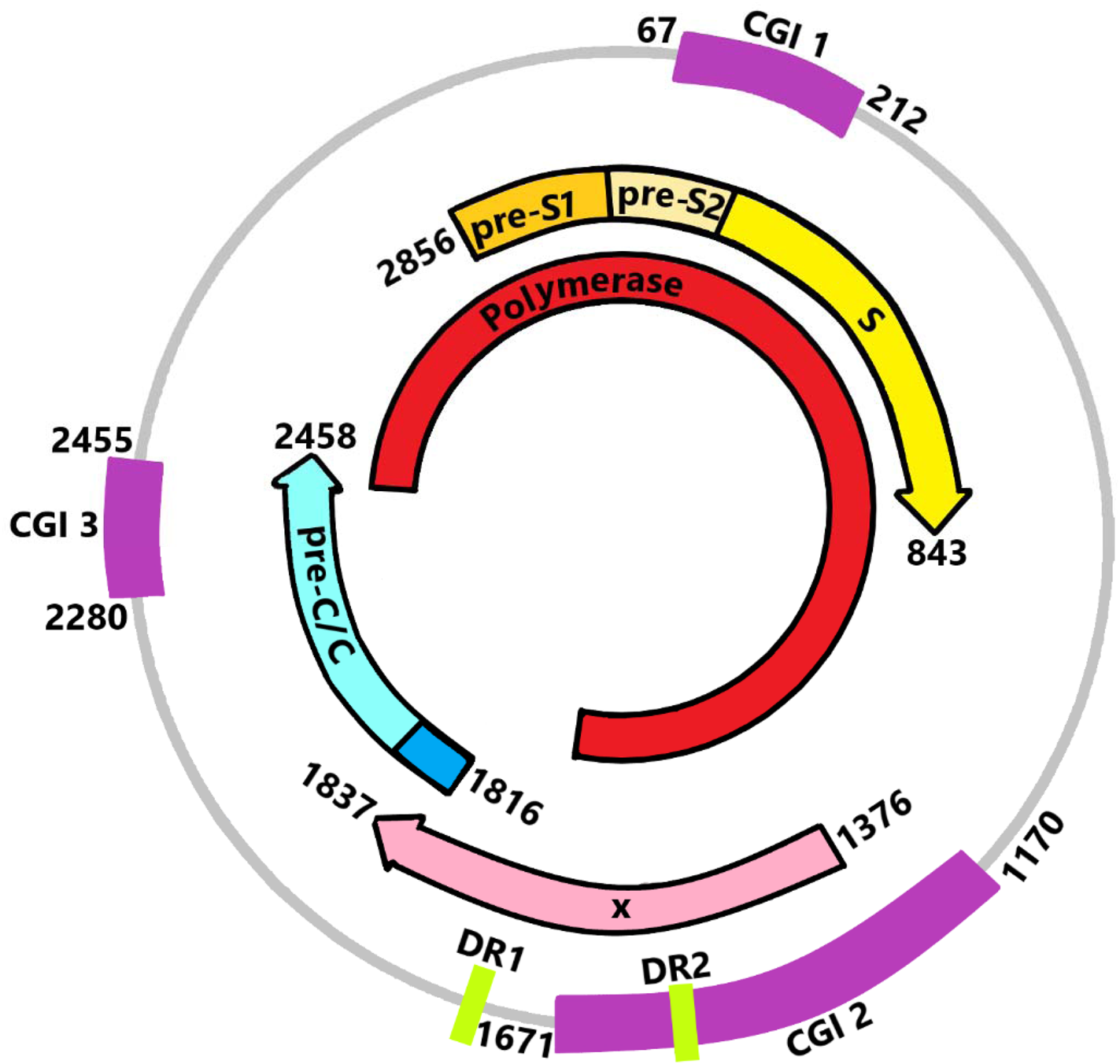
Fig. 1. DNA, transcripts and location of CpG islands (CGIs) of HBV. CGI 1 is located at the start site of the S region around nucleotide 67 to nucleotide 212, while CGI 2 (nucleotide 1170 to nucleotide 1671) is located between the enhancer I and the promoter of the X gene. CGI 3 (nucleotide 2280 to nucleotide 2455) comprises the start site of the P region with partial coverage of the C gene.
HBV has a unique replication strategy that allows the virus to persist in the hepatocytes of the host. The persistence of HBV infection is maintained by HBV cccDNA in the nuclei of hepatocytes. HBV cccDNA associate with histones and HBc to form a stable minichromosome in the nucleus of the hepatocyte (Refs Reference Caballero28, Reference Chong30, Reference Levrero31). Following infection, HBV attaches to the surface of hepatocyte, releasing its viral nucleocapsid into cytoplasm. The viral capsid then attaches to nucleus, releasing its partial double-stranded DNA into nucleus. The rcDNA is converted to cccDNA by the host repair machineries. (Refs Reference Seeger and Mason29, Reference Mitra32, Reference Wei and Ploss33). HBV cccDNA is an important replicative intermediate for the transcription of viral RNAs including pregenomice RNA (pgRNA) by cellular RNA polymerase. The pgRNA is packaged into viral capsids and is reverse transcribed into new rcDNA. The newly formed rcDNA could enter the nucleus again to form cccDNA. Else, it would be released as new progeny virion after encapsidation and envelopment (Fig. 2) (Refs Reference Belloni34–Reference Zhang38).

Fig. 2. HBV DNA replication in hepatocyte.
Influences of HBV genotypes, viral load and genome mutations on HBV-related HCC development
To date, a total of 10 genotypes and several subtypes of HBV have been classified with different geographic distributions (Refs Reference Velkov39, Reference Lin and Kao40) (Table 1). Liver cirrhosis may be the main factor for the development of HCC in CHB patients. However, HBV genotypes, viral load, genomic mutations and integration of HBV DNA are viral factors that have been associated with the severity of liver injury and the development of HCC (Refs Reference An8, Reference Lin and Kao40–Reference Tu42).
Table 1. Geographical distribution of HBV genotypes

HBV genotypes may influence the risk of HCC development in each country. Genotypes B and C are the predominant genotypes in Asia (Ref. Reference Velkov39). Two meta-analysis studies showed that patients with genotype C are usually at higher risk of developing HCC than those with genotype B (Refs Reference Wong43, Reference Raffetti, Fattovich and Donato44). On the other hand, it has been reported that the rate of HCC development is higher in patients with genotypes A, C and F compared with genotypes B and D in the Alaska native population (Ref. Reference Ching45) and children and young adults with genotype F1 are also at higher risk of developing HCC (Ref. Reference Gounder46).
HBV viral load is an independent risk factor for liver disease progression. Persistently high viral replication, as indicated by high circulating HBV DNA levels, increases the risk of developing severe liver disease (Ref. Reference Yang47). Patients with HBV genotype C have a higher viral load compared with patients with genotype B (Ref. Reference Raffetti, Fattovich and Donato44). This finding is consistent with the in vitro study which showed that the cell line exposed to genotype C had a higher HBV DNA level than that exposed to genotype B and the HBV DNA level in cells exposed to genotype D was higher than that exposed to genotype A (Ref. Reference Sugiyama48).
There is increasing evidence that the specific nucleotide-level mutations in the regulatory regions of HBV DNA may contribute to HCC. It has been reported that mutations in the PreC/C, PreS/S, X and P regions accelerate progression to HCC in CHB patients (Refs Reference Yang49, Reference Yang50). The most frequent mutations were identified in the PreC/C region, including various basal core promoter mutations (A1762T/G1764A, T1753V, C1766T and T1768A), C1653T mutation in the enhancer region II and G1896A mutation in the PreC region (Refs Reference Rajoriya41, Reference Yang50–Reference Wahyuni52). Based on a database analysis of 340 C1 strains from GenBank, approximately 95% of HCC patients with genotype C1 have mutations in the basal core promoter region, and this is in concordance with the high frequency of these mutations in genotype C1 observed in Cambodian CHB cases. Therefore, the authors concluded that the mutations in genotype C1 are highly correlated with the development of HCC. (Ref. Reference Chuon51).
HBV DNA integration is known to be a factor in the development of HCC, although the mechanisms of integration are not well understood (Refs Reference Sung53, Reference Yang54). Host genome modifications by integrated HBV DNA, including genomic instability, cis-activation of tumour-promoting genes and expression of mutant HBV protein, have been associated with HCC progression (Refs Reference Dandri9, Reference Tu42). Tu et al. reported in their cell line study that HBV DNA integration probably occurs in the early phase of HBV infection (Ref. Reference Tu55). This finding is contrary to the result of another study that indicated that HBV DNA integration does not occur only in the early stage and that HCC patients have a higher integration rate than CHB patients (Ref. Reference Yang54).
CpG Islands of HBV DNA
Cytosine-phosphate-guanine is abbreviated as CpG. CpG islands (CGIs) are regions of the genome rich in cytosine-guanine dinucleotide repeats, where the guanine is at the 3′ end of cytosine on a linear sequence (CpG) (Refs Reference Charostad18, Reference Jones56, Reference Moore, Le and Fan57). In the HBV genome, CGIs 1, 2 and 3 are present together in most HBV genotypes (Table 2). Some may contain only 2 CGIs, and the distribution of CGIs is not HBV genotype-specific (Refs Reference Hou58–Reference Zhang60). CGI 2 and 3 are more conserved compared to CGI 1 (Refs Reference Xue59, Reference Guo61, Reference Zhong62). The absence of CGI 1 was observed in the HBV genome with lower density in cytosine-guanine dinucleotide repeats (Ref. Reference Hou58). The loss of CGI 1 is more frequent in genotype C (approximately 67–80%) compared to genotype B (Refs Reference Xue59, Reference Zhang60). These distribution characteristics of CGIs might be associated with different HBV disease courses. Hou et al. (Ref. Reference Hou58) and Xue et al. (Ref. Reference Xue59) speculated that the absence of CGI 1 in genotype C may promote severe liver disease, which is related to the methylation status of CGIs. CGIs are located within the promoter regions of HBV genes (Fig. 1). CpG methylation involves the addition of a methyl group to the fifth carbon of cytosine residues to form 5-methylcytosine (Refs Reference Charostad18, Reference Jones56, Reference Moore, Le and Fan57). In general, CpG methylation in regulatory regions reduces gene expression. Therefore, it is conceivable that CpG methylation in CGIs may inhibit viral gene expression because these CGIs fall within the gene regulatory region, highlighting their potential role in regulating the viral life cycle and disease pathogenesis.
Table 2. The location and size of the three conventional CpG Islands within typical HBV genomes of different genotypes

bp, base pair; CGI, CpG island.
Effects of DNA methylation in CGI regions by DNMTs
DNMTs are host enzymes of mammals, consisting of DNMT1, DNMT2, DNMT3A, DNMT3B and DNMT3L. DNMT1, DNMT3A and DNMT3B are members of the family of canonical enzymes involved in DNA methylation of CGIs (Refs Reference Lyko27, Reference Jones56). Approximately 70% of gene promoters are located within CGI regions, and transcriptional factors (TFs) initiate gene expression via a CG-rich promoter sequence (Refs Reference Héberlé and Bardet63, Reference Lambert64). Methylation in CGI inhibits transcription initiation by interfering with the binding of TFs, thereby repressing gene expression (Refs Reference Lyko27, Reference Moore, Le and Fan57, Reference Héberlé and Bardet63, Reference Koumbi and Karayiannis65). The interference with binding may be due to the binding sites being rendered incompatible for TFs by methylated cytosine or due to repressor proteins binding to the methylated cytosine sites, thereby rendering the binding sites inaccessible for binding by TFs (Fig. 3). Three types of repressor proteins, namely methyl-CpG-binding domain (MBD) protein, ubiquitin-like containing plant homeodomain and RING finger domain (UHRF) protein, and zinc-finger protein are known methyl-binding proteins (MBPs) that work collectively to exert such an inhibitory effect (Refs Reference Moore, Le and Fan57, Reference Hoelzer, Shackelton and Parrish66, Reference Kim67). In contrast, several studies expounded the opposite view that the methylation of DNA may activate gene transcription. The activation is probably because methylated sites are recognised by TFs or the methylated site is located within a negative regulatory region of the gene (Refs Reference Lyko27, Reference Héberlé and Bardet63, Reference Yin68). The precise functions of DNA methylation await further investigation.

Fig. 3. Actions of methylation on transcriptional start site (TSS) of a gene. CpG island (GCI) in the TSS is methylated by DNA methyltransferases (DNMTs). The binding sites is being rendered incompatible for transcriptional factors (TFs) due to methylated cytosine or repressor proteins binding to the methylated cytosine sites, thereby repressing gene transcription.
Up-regulation of DNMTs activity after HBV infection
Several studies reported that host DNMT activity is upregulated during HBV infection (Refs Reference Park26, Reference Brezgin69–Reference Zeisel, Guerrieri and Levrero71). The increased production of DNMTs is believed to be due to the recognition of integrated HBV DNA as a foreign entity in infected hepatocytes (Refs Reference Dandri9, Reference Koumbi and Karayiannis65, Reference Hoelzer, Shackelton and Parrish66, Reference Tian72, Reference Watanabe73). On the other hand, the upregulation of DNMT activity could be activated by HBV proteins (Refs Reference Charostad18, Reference Zeisel, Guerrieri and Levrero71). The resulting methylation has been viewed as an attempt by the host to prevent transcription of viral genes, and by HBV to repress the expression of TSGs or simultaneously manipulate the expression of immunoregulatory genes (Refs Reference Dandri9, Reference Brezgin69).
Host cell DNMTs may methylate promoter regions of HBV DNA such as CGI 1 and CGI 2, to inhibit viral gene expression (Refs Reference Jain21, Reference Kaur22, Reference Kim67, Reference Vivekanandan70, Reference Wang, Wang and Wang74). This was demonstrated in a study in which upregulation of host DNMTs by overexpression of microRNA-20a (miR-20a) suppressed HBV replication via the induction of DNA methylation of cccDNA (Refs Reference Moon75, Reference Park76). It was postulated that miR-20a facilitates the binding of Argonaute2 (AGO2), an effector of RNA-induced silence complex, to cccDNA and this event induces HBV DNA methylation in human hepatoma cells to suppress gene transcription and thus HBV replication (Ref. Reference Moon75). On the other hand, another study demonstrated that the methylation of C1619 within the CGI of cccDNA induces viral replication (Ref. Reference Lee25). C1619 is localised within the negative regulatory element (NRE) of the HBc promoter region. The NRE has a downregulatory role in HBc gene expression (Ref. Reference Sun77). This is consistent with a DNMT1 knockout study showing that unmethylated C1619 diminishes HBV replication and transcription (Refs Reference Lee25, Reference Guo61). Although the upregulation of DNMTs activity affects HBV replication, a study using an HBV-expressing cell line demonstrated an increase in cccDNA levels and suggested that the silencing of APO3a or APO3b (effector proteins that eliminate cccDNA) induces cccDNA accumulation (Ref. Reference Brezgin69).
In parallel, DNMTs also methylate TSGs (Refs Reference Zhong62, Reference Vivekanandan70). Methylated TSGs, such as the Adenomatous Polyposis Coli, Ras Association Domain Family 1 isoform A, and Glutathione S-transferase Pi 1, have been detected in serum or biopsy samples from HCC patients (Refs Reference Dong19, Reference Jain21, Reference Ye23), as well as in cell lines infected with HBV (Ref. Reference Vivekanandan70). These observations led researchers to conclude that methylation occurs in both viral and host genes upon HBV infection. Furthermore, host DNMTs simultaneously alter host DNA damage by downregulating DNA damage repair factor expression, as the promoter of DNA damage repair genes is methylated due to DNMT overexpression (Ref. Reference Brezgin69).
The multitasking role of HBx protein in DNA methylation and HBV DNA replication
The upregulation of host DNMTs activity by HBx protein has been demonstrated in many studies. Highly expressed HBx protein enhances methylation activities in the liver cells (Refs Reference Lee25, Reference Park26). As a result, DNMT methylates host TSGs and immunoregulatory genes and may collaterally methylate HBV DNA (Ref. Reference Vivekanandan70). HBx acts as a transactivator for host DNMT gene expression (Ref. Reference An8). This protein could activate host DNMT transcriptional activity by promoting histone acetylation in the promoter region of canonical DNMT genes except for DNMT3b (Ref. Reference Park26).
HBx protein dysfunction may also decrease viral DNA level and HBV transcripts production (Refs Reference Chou78, Reference Lucifora79). This HBV protein has a unique function in regulating viral DNA replication in several ways. In hepatocytes, HBx increases acetylation of the cccDNA minichromosome. This leads to the broadening of nucleosome spacing and release of the HBV genome for transcriptional activity (Refs Reference Lucifora79–Reference Pollicino81). Simultaneously, HBx induces the recruitment of transcriptional factors and co-activators to the cccDNA minichromosome to further activate the transcriptional activity of HBV DNA (Ref. Reference Wang, Wang and Wang74). It has been shown to protect pgRNA from binding several host miRNAs including miR-224, miR-137 and miR-596, which may inhibit viral transcription. By targeting the promoter region of miRNAs, it downregulates their expression (Ref. Reference Guerrieri82). HBx also promotes the synthesis of deoxynucleoside triphosphates for HBV DNA replication by activating the R 2 subunit of ribonucleotide reductase (Refs Reference Cohen83, Reference Ricardo-Lax84). The inhibitory effects of the Smc5/6 complex for structural maintenance of host chromosomes are abrogated by HBx to upregulate viral transcriptional activity. This protein promotes degradation of the Smc5/6 complex through interaction with the host DNA damage binding protein 1 (Refs Reference Wang, Wang and Wang74, Reference Murphy85, Reference Sekiba86). These diverse roles of HBx provide a new perspective in HBV methylation, transcription and replication.
HBV DNA methylation patterns
Three common CGIs of HBV DNA were studied to understand their methylation patterns during CHB-related HCC development (Table 3). Vivekanandan et al. (Ref. Reference Vivekanandan, Thomas and Torbenson87) showed that CGI 1 was more methylated than CGI 2 in both non-HCC and HCC tissues. Kaur et al. (Ref. Reference Kaur22) found that occult HCC and cirrhotic samples had higher methylation of CGI 1 than CGI 2 and 3, and that hypermethylation of CGI 1 was strongly associated with liver cancer development. A similar finding for CGI 1 was demonstrated in another study. In that study, it was found that CGI 1 had a higher level of methylation in HCC and CHB cases than cirrhosis cases (Ref. Reference Jain21). In contrast, another study showed that CGI 1 was rarely methylated in the early stage of CHB infection, but patients with CHB-related liver fibrosis exhibited a higher CGI 1 methylation level (Ref. Reference Zhang88). From this finding, it could be postulated that HBV DNA methylation is associated with severe HBV disease courses such as liver fibrosis.
Table 3. The HBV DNA methylation patterns

CGI, CpG island, CHB, chronic hepatitis B, HCC, hepatocellular carcinoma.
An interesting observation is that CGI 2 methylation is minimal in HCC and CHB cases (Refs Reference Jain21, Reference Kaur22, Reference Vivekanandan, Thomas and Torbenson87). This low level of methylation could be explained by the postulation that in HCC, the activation of viral replication requires HBc protein to rebind to CGI 2, and this binding may prevent the methylation of CGI 2 (Refs Reference Guo61, Reference Chong89). Consequently, a lower level of methylation has been observed in CGI 2 (Refs Reference Jain21, Reference Kaur22). This observation is in contrast to another study in which Zhang et al. reported that CGI 2 was highly methylated in hepatitis samples, and that methylation of CGI 2 was correlated with lower expression of HBs protein (Ref. Reference Zhang88). Unexpectedly, non-CpG methylation was found in CGI 2 of cccDNA, which was attributed to de novo methylation (Ref. Reference Jain21). Jain et al. (Ref. Reference Jain21) observed that CGI 3 was the most methylated among the three CGIs in their HCC samples. Approximately 53% of the cytosine-guanine dinucleotide repeats were methylated in CGI 3. Zhang et al. (Ref. Reference Zhang88) recorded the same observation in their samples from CHB patients, but additionally found that higher level methylation of CGI 3 impaired HBs protein production.
There is a strong relationship between HBV DNA methylation and HCC development, but the exact underlying mechanism has not been fully elucidated. A lower degree of CGIs methylation has been observed in CHB or cirrhotic cases than in HCC cases (Refs Reference Fernandez20–Reference Kaur22) but hypermethylation of CGI 1 and CGI 3 in HBV cccDNA occurs during hepatocarcinogenesis and the methylation status of CGI 1 is correlated with CGI 3 in HBV-related HCC (Ref. Reference Jain21). The methylation level of CGI 2 has been reported to be minimal among the three CGIs (Refs Reference Jain21, Reference Kaur22, Reference Vivekanandan, Thomas and Torbenson87). This could be related to the situation of CGI 2 within the regulatory region for several HBV mRNA transcripts (Refs Reference Jain21, Reference Guo61, Reference Zhang88) and unmethylated CGI 2 being required for viral transcription in the host cells. It has been noted that methylated CGI 2 significantly reduces the amount of pgRNA and HBV proteins (Ref. Reference Vivekanandan, Thomas and Torbenson90). The methylation status of CGIs is associated with the production of HBV proteins in hepatocytes such as HBs and HBx. The role of HBx is significant in CHB infection. It has been shown to induce the upregulation of DNMTs activity in hepatocytes (Refs Reference Lee25, Reference Park26, Reference Vivekanandan70). Taken together, it appears plausible that high level methylation of host TSGs and CGIs of HBV DNA, especially in CGIs 1 and 3, along with the high level of pgRNA expression, allows HBV to persist in the host and leads to the development of HCC (Refs Reference Jain21, Reference Vivekanandan, Thomas and Torbenson90).
The methylation status of CGIs in cccDNA has been shown to correlate with the stage of liver fibrosis (Refs Reference Kim67, Reference Zhang88). As liver fibrosis often progresses to cirrhosis and HCC (Refs Reference Guan and Lui7, Reference An8), it is reasonable to link cccDNA methylation to the development of HCC. However, the accumulation of methylated cccDNA is also associated with patient age and duration of HBV infection (Refs Reference Kim67, Reference Zhang88), suggesting that the methylation level of cccDNA may be a consequence of prolonged CHB infection and not a reflection of HCC development. In a recent study on the kinetics of decay, cccDNA mutated by lamivudine resistance in the treated HBV patients was used to indicate the half-life of cccDNA. The authors reported that the half-life of cccDNA in human’ hepatocytes is several months rather than years (Ref. Reference Huang91). This result indicated that the duration of CHB infection may not be directly related to the accumulation of methylated cccDNA in hepatocytes. On the other hand, cccDNA methylation is related to the dynamics of immune clearance in CHB-infected patients, as the degree of CGI 2 methylation is much higher in HBeAg-negative patients than in HBeAg-positive patients (Refs Reference Zhang88, Reference Guo92). This observation is consistent with a HepG2 cell line study, in which a high-level methylation of CGI 2 was reported to decrease the production of intercellular HBe protein (Ref. Reference Nakamura93). Moreover, methylation in CGI 2 and/or 1 may down-regulate the expression of hepatitis B surface (HBs) protein (Refs Reference Vivekanandan, Thomas and Torbenson90, Reference Nakamura93, Reference Graumann94). The role of CGI methylation in HCC remains controversial. Further studies on this topic are needed for a better understanding of the contribution of cccDNA methylation to the development of HCC.
Methylated cccDNA as the potential biomarker for early HCC
Biomarkers are useful if they are applicable for early detection, monitoring, prediction and prognosis of disease. In this context, they must be reliably detectable or measurable in patients' specimens. Current biomarkers for HCC diagnosis in blood are AFP, AFP-L3 and DCP (Refs Reference Choi95–Reference Zacharakis, Aleid and Aldossari99). These biomarkers suffer from low sensitivity. At the cut-off value of 30 ng/ml AFP, sensitivity and specificity are 65 and 89%, respectively (Ref. Reference Zhou, Liu and Luo100). Only 10–20% of early-stage small HCC could be detected with AFP (Ref. Reference Zhou, Liu and Luo100). With a cut-off of 35%, AFP-L3 has 100% specificity for early diagnosis. Overall, 35% of patients with small HCC (<3 cm) could be detected with this cut-off. In this case, the sensitivity of AFP-L3 decreases to 33% (Refs Reference Zhao, Ju and Li14, Reference Leerapun96, Reference Zacharakis, Aleid and Aldossari99–Reference Debruyne and Delanghe101). A combination of AFP, AFP-L3 and DCP is only slightly more sensitive than using any one of them alone in HCC diagnosis (Refs Reference Parikh17, Reference Choi95, Reference Zacharakis, Aleid and Aldossari99, Reference Marrero102). Therefore, reliable biomarkers for the diagnosis of early-stage HCC are currently scarce.
Following the recent discovery of HBV DNA methylation in CHB- and HBV-related HCC patients (Refs Reference Jain21, Reference Kaur22, Reference Zhong62, Reference Park76, Reference Zhang88, Reference Guo92) and the detectability of HBV cccDNA in patient's serum or liver biopsy samples (Refs Reference Huang103–Reference Wong109), the new challenge of using this phenomenon for the detection of early-stage HCC is opening up. HBV DNA is methylated during the upregulation of DNMTs activity in hepatocyte upon HBV infection. The cccDNA is a replicative intermediate in the nuclei of hepatocytes, but it may be released from hepatocytes into blood circulation upon liver injury or other less well-known secretory mechanisms (Refs Reference Huang103, Reference Yuen108).
Serum cccDNA level was positively correlated with intrahepatic cccDNA level, suggesting that cccDNAs present in serum are likely released from infected hepatocytes (Refs Reference Huang103, Reference Minosse105, Reference Takkenberg110, Reference Cabrerizo111). This is consistent with the results of an in vitro study in which a high amount of cccDNA is released into the culture medium with hepatic cell death and the cccDNA is detected in both intracellular and extracellular environments (Ref. Reference Liu112). The intrahepatic cccDNA level in tumour tissue is higher than in non-tumour tissue (Ref. Reference Wong109). The fact that HBV cccDNA could effectively predict HCC recurrence, supporting the use of HBV cccDNA as an early biomarker for HCC (Ref. Reference Hsu113). In a recent study, cccDNA was detected in nearly 90% of serum from HCC patients, compared with 53% in non-HCC patients (Ref. Reference Huang103).
It has been observed that HCC patients have higher serum cccDNA levels compared with non-HCC patients; therefore, serum cccDNA levels may be used to distinguish between HCC and non-HCC patients. The combination of serum cccDNA and HBV DNA levels has a specificity of 93.7% and a sensitivity of 74.5% for early HCC diagnosis (Ref. Reference Huang103). The degree and number of HBV DNA methylation events were observed more frequently in HCC patients compared with patients without HCC (Ref. Reference Jain21). Unlike serum AFP, the measurement of cccDNA is unlikely to be affected by normal physiological factors such as pregnancy or liver inflammation (Ref. Reference Zhao, Ju and Li14). Serum cccDNA may be a better biomarker of liver damage because it can be detected much earlier than serum alanine aminotransferase (ALT) levels in CHB patients. Serum cccDNA level rose earlier than ALT flare. However, cccDNA was progressively reduced as the ALT level increased (Ref. Reference Chen, Sze and He114). Therefore, it is conceivable that circulating methylated cccDNAs have great potential as an early biomarker for HCC. Further studies on circulating methylated cccDNAs are required to better define its role in HCC diagnosis.
Conclusion and perspective
Following HBV infection, the expression of host DNMTs is upregulated, resulting in methylation of both host and viral genes. HBV cccDNA methylation involving CGIs affects viral replication and is associated with progression to HCC. HBV DNA methylation is one of the side effects induced by overexpression of DNMTs in the liver of patients. Other effects such as the inactivation of TSGs and host genome damage have been reported to be associated with DNMT activity in HBV infection. Thus, with the involvement of many factors such as the accumulation of methylated cccDNA, HBV DNA integration, and HBV genome mutations still unresolved, the role of HBV DNA methylation is still contentious. Detectable methylated HBV cccDNA in human serum makes it a suitable and less invasive biomarker for early detection of HCC in CHB patients. While HBV cccDNA methylation would be useful for early detection of HBV-related HCC, the challenges are to simplify the laborious methods of viral cccDNA isolation and to develop robust detection methods for cccDNA methylation. A sensitive, high-throughput method is important for the detection of cccDNA. Unfortunately, many uncertainties remain regarding the mechanisms and significance of HBV DNA methylation, further explorations are necessary for clarification.
Acknowledgements
Not applicable.
Author contributions
Conceptualisation, J.B.C.; Writing – original draft preparation, W.F.L.; Writing – review and editing, Y.F.N, J.B.C., K.K.T, S.K.O, S.C.P, J.J.B and R.M.; Funding acquisition, J.B.C. All authors approved the final manuscript.
Financial support
This work was supported by Ministry of Higher Education Malaysia, grant number is FRGS/1/2019/SKK08/SYUC/03/2 and Sunway Medical Centre research grant, grant number is SRC/006/2017/FR.
Conflict of interest
The authors declare no conflict of interest.









